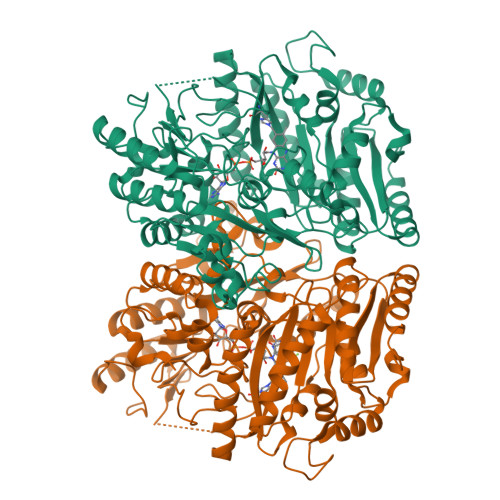
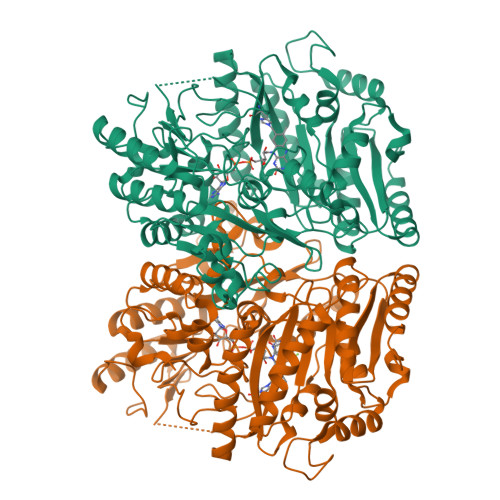
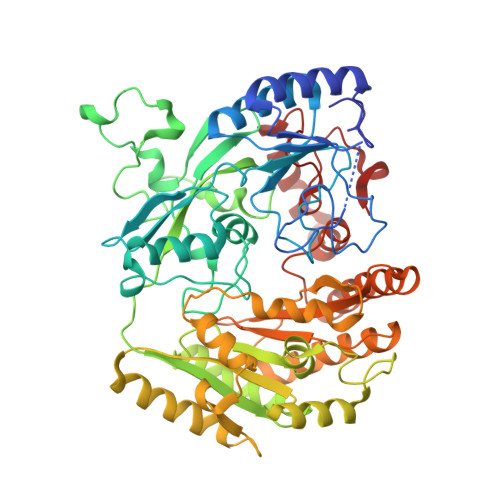
Inversion of Stereospecificity in Vanillyl-Alcohol Oxidase
Van Der Heuvel, R.H.H., Fraaije, M.W., Espinosa, M.F., Mattevi, A., Van Berkel, W.J.H.(2000) Proc Natl Acad Sci U S A 97: 9455
- PubMed: 10920192
- DOI: https://doi.org/10.1073/pnas.160175897
- Primary Citation of Related Structures:
1E0Y - PubMed Abstract:
Vanillyl-alcohol oxidase (VAO) is the prototype of a newly recognized family of structurally related oxidoreductases sharing a conserved FAD-binding domain. The active site of VAO is formed by a cavity where the enzyme is able to catalyze many reactions with phenolic substrates. Among these reactions is the stereospecific hydroxylation of 4-ethylphenol-forming (R)-1-(4'-hydroxyphenyl)ethanol. During this conversion, Asp-170 is probably critical for the hydration of the initially formed p-quinone methide intermediate. By site-directed mutagenesis, the putative active site base has been relocated to the opposite face of the active site cavity. In this way, a change in stereospecificity has been achieved. Like native VAO, the single mutants T457E, D170A, and D170S preferentially converted 4-ethylphenol to the (R)-enantiomer of 1-(4'-hydroxyphenyl)ethanol. The double mutants D170A/T457E and D170S/T457E exhibited an inverted stereospecificity with 4-ethylphenol. Particularly, D170S/T457E was strongly (S)-selective, with an enantiomeric excess of 80%. The crystal structure of D170S/T457E, in complex with trifluoromethylphenol, showed a highly conserved mode of ligand binding and revealed that the distinctive catalytic properties of this mutant are not caused by major structural changes.
Organizational Affiliation:
Department of Biomolecular Sciences, Laboratory of Biochemistry, Wageningen University, The Netherlands.