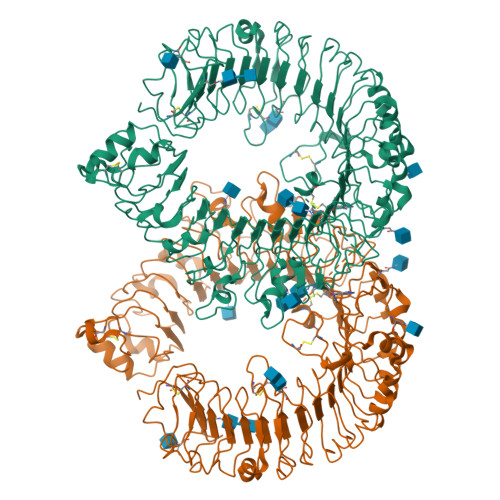
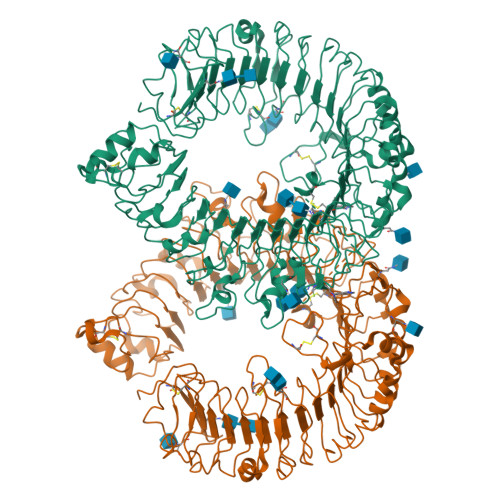
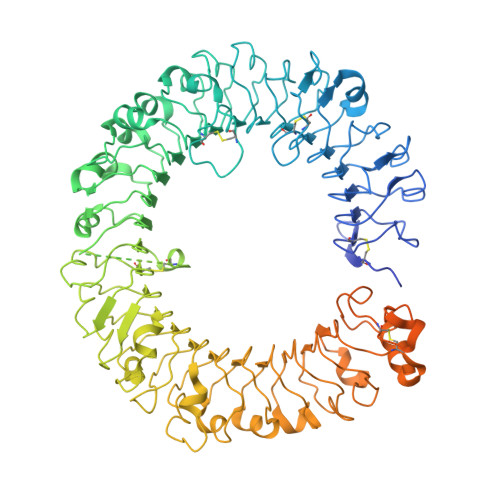
A novel Toll-like receptor 7/8-specific antagonist E6742 ameliorates clinically relevant disease parameters in murine models of lupus.
Ishizaka, S.T., Hawkins, L., Chen, Q., Tago, F., Yagi, T., Sakaniwa, K., Zhang, Z., Shimizu, T., Shirato, M.(2023) Eur J Pharmacol 957: 175962-175962
- PubMed: 37544422
- DOI: https://doi.org/10.1016/j.ejphar.2023.175962
- Primary Citation of Related Structures:
7YTP, 7YTX - PubMed Abstract:
The sensing of self RNA by the endosomal Toll-like receptors (TLRs) 7 and 8 initiates pathogenic mechanisms underlying the autoimmune disease lupus. A blockade of the TLR7/8 signals may, therefore, be a novel therapeutic intervention for lupus. To test the hypothesis, a novel compound E6742 that blocks TLR7/8 activation was identified. The mode of action of E6742 was investigated by analysis of the tertiary structure of TLR7 and 8 in complex with E6742. The in vitro activities of the compound were examined in cellular systems and its therapeutic potential was evaluated in murine lupus models. Tertiary structures of the extracellular domain of TLR7 and 8 in complex with E6742 showed that E6742 binds specifically and non-covalently to the hydrophobic pocket located at the interface of TLR7 or TLR8 homodimers. E6742 potently and selectively inhibited several TLR7/8-mediated cytokine responses in human PBMC. In two mouse models of lupus, oral dosing of E6742 after the onset of disease suppressed increase in autoantibodies and blocked the advance of organ damage. Collectively, the data show that TLR7/8 activation contributes to disease progression and its blocking by E6742 has potential as a therapeutic intervention for lupus.
Organizational Affiliation:
Eisai Inc., Eisai Center for Genetics Guided Dementia Discovery, MA, USA.