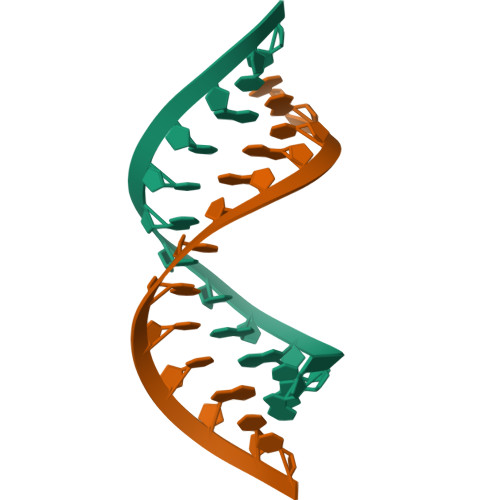
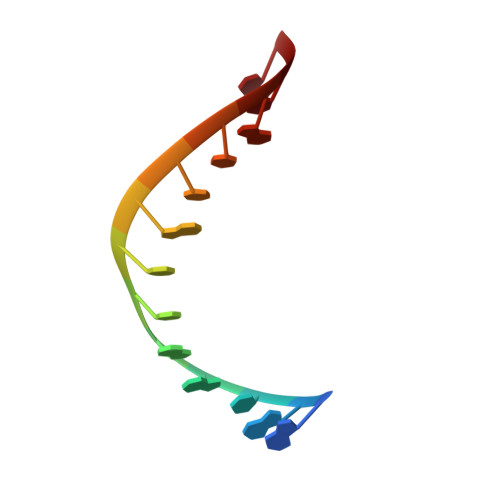
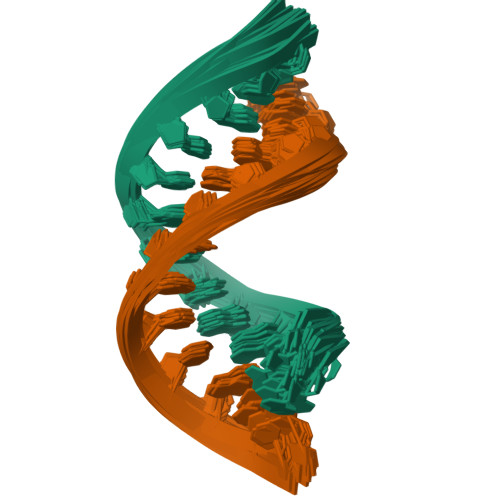
Structures of small molecules bound to RNA repeat expansions that cause Huntington's disease-like 2 and myotonic dystrophy type 1.
Chen, J.L., Taghavi, A., Frank, A.J., Fountain, M.A., Choudhary, S., Roy, S., Childs-Disney, J.L., Disney, M.D.(2024) Bioorg Med Chem Lett : 129888-129888
- PubMed: 39002937
- DOI: https://doi.org/10.1016/j.bmcl.2024.129888
- Primary Citation of Related Structures:
9CPD, 9CPG, 9CPI, 9CPJ - PubMed Abstract:
Trinucleotide repeat expansions fold into long, stable hairpins and cause a variety of incurable RNA gain-of-function diseases such as Huntington's disease, the myotonic dystrophies, and spinocerebellar ataxias. One approach for treating these diseases is to bind small molecules to these structured RNAs. Both Huntington's disease-like 2 (HDL2) and myotonic dystrophy type 1 (DM1) are caused by a r(CUG) repeat expansion, or r(CUG) exp . The RNA folds into a hairpin structure with a periodic array of 1 × 1 nucleotide UU loops (5'CUG/3'GUC; where the underlined nucleotides indicate the Us in the internal loop) that sequester various RNA-binding proteins (RBPs) and hence the source of its gain-of-function. Here, we report nuclear magnetic resonance (NMR)-refined structures of single 5'CUG/3'GUC motifs in complex with three different small molecules, a di-guandinobenzoate (1), a derivative of 1 where the guanidino groups have been exchanged for imidazole (2), and a quinoline with improved drug-like properties (3). These structures were determined using NMR spectroscopy and simulated annealing with restrained molecular dynamics (MD). Compounds 1, 2, and 3 formed stacking and hydrogen bonding interactions with the 5'CUG/3'GUC motif. Compound 3 also formed van der Waals interactions with the internal loop. The global structure of each RNA-small molecule complexes retains an A-form conformation, while the internal loops are still dynamic but to a lesser extent compared to the unbound form. These results aid our understanding of ligand-RNA interactions and enable structure-based design of small molecules with improved binding affinity for and biological activity against r(CUG) exp . As the first ever reported structures of a r(CUG) repeat bound to ligands, these structures can enable virtual screening campaigns combined with machine learning assisted de novo design.
Organizational Affiliation:
Department of Chemistry, The Scripps Research Institute, 130 Scripps Way, Jupiter, FL 33458, USA; Department of Biochemistry and Biophysics, Center for RNA Biology, University of Rochester Medical Center, Rochester, NY 14642, USA.