Full Text |
QUERY: Gene Name = "XRCC6" | MyPDB Login | Search API |
| Search Summary | This query matches 32 Structures. |
Structure Determination MethodologyScientific Name of Source OrganismTaxonomyExperimental MethodPolymer Entity TypeRefinement Resolution (Å)Release DateEnzyme Classification NameSymmetry Type | 1 to 25 of 32 Structures Page 1 of 2 Sort by
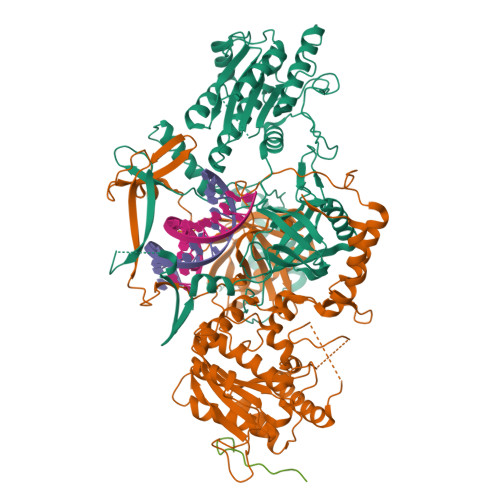
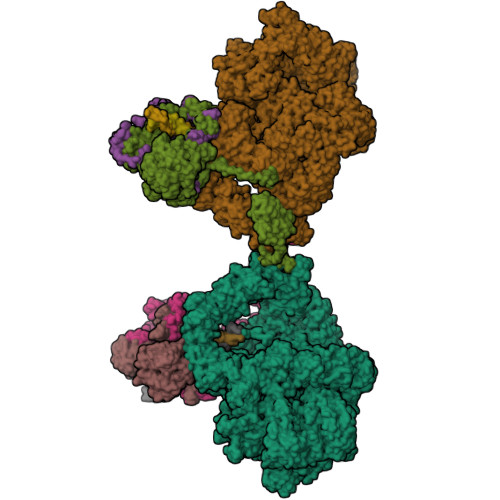
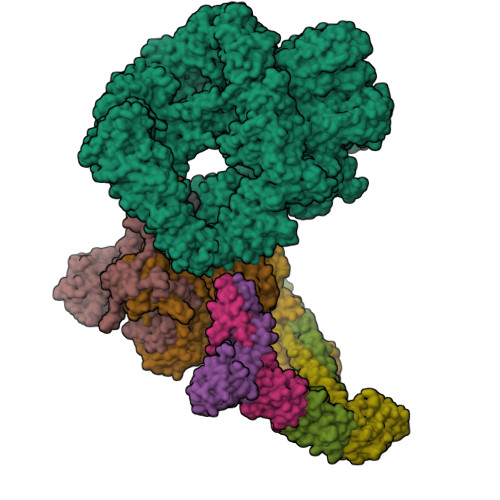
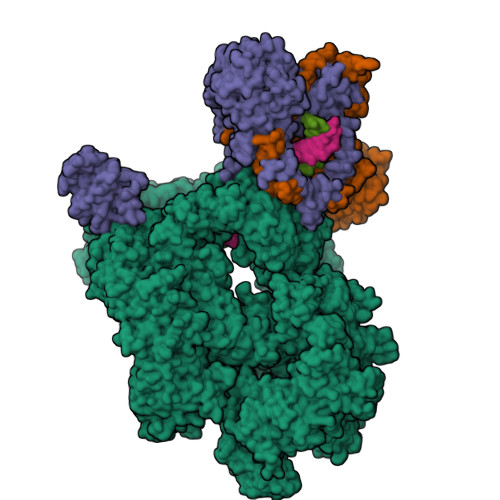
Cryo-EM structure of Human DNA-PK HoloenzymeYin, X., Liu, M., Tian, Y., Wang, J., Xu, Y. (2017) Cell Res 27: 1341-1350
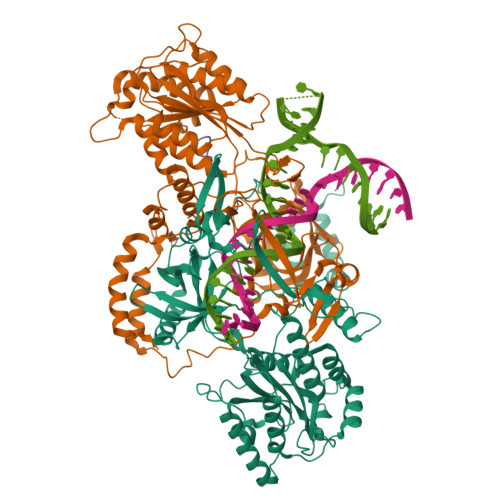
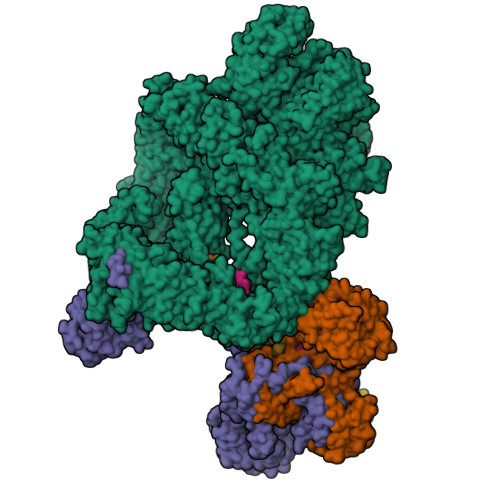
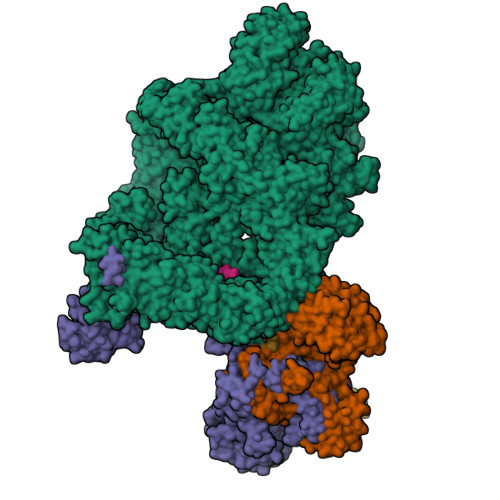
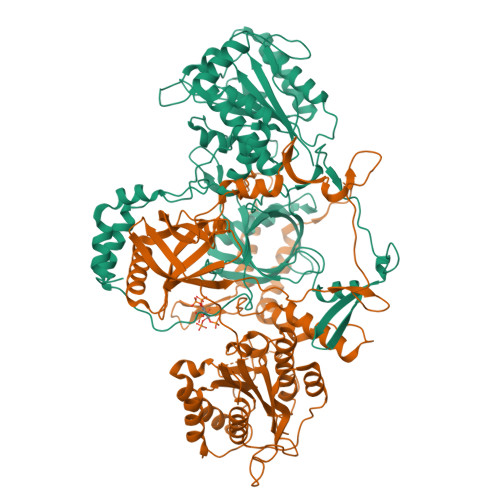
Complex of APLF factor and Ku heterodimer bound to DNANemoz, C., Legrand, P., Ropars, V., Charbonnier, J.B. (2018) Nat Struct Mol Biol 25: 971-980
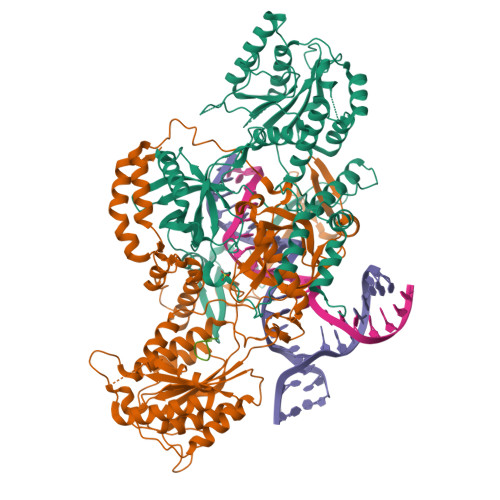
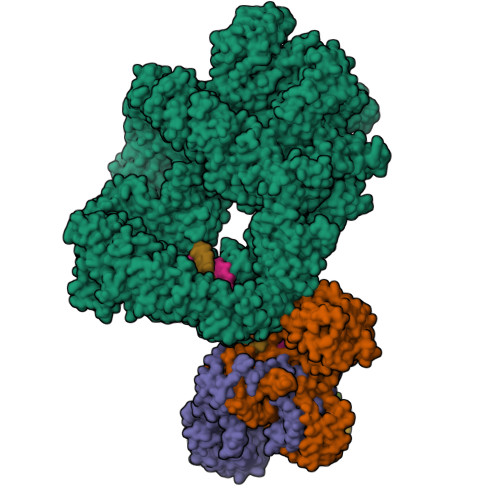
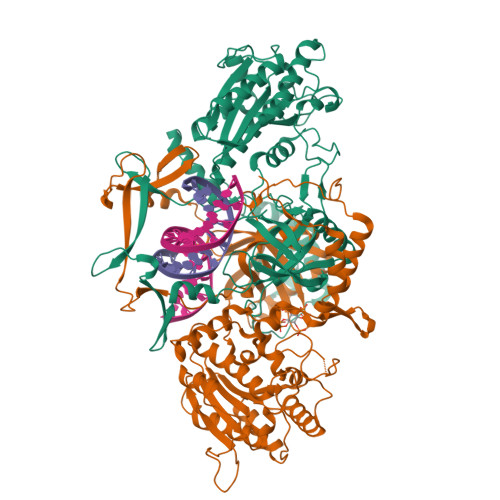
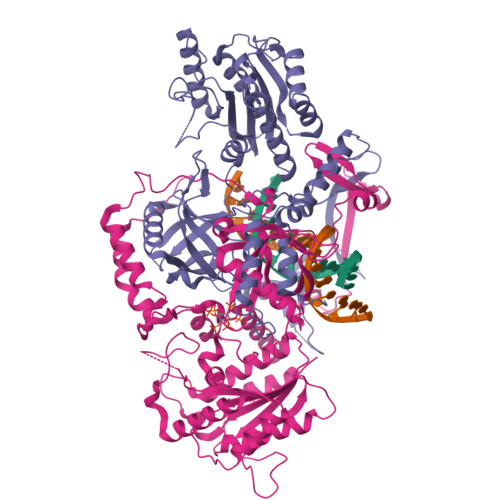
Complex of XLF and heterodimer Ku bound to DNANemoz, C., Legrand, P., Ropars, V., Charbonnier, J.B. (2018) Nat Struct Mol Biol 25: 971-980
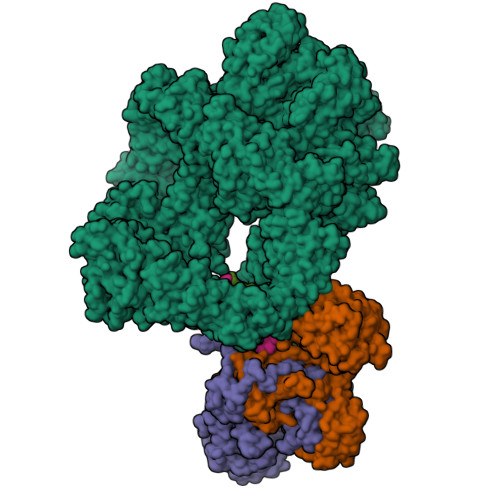
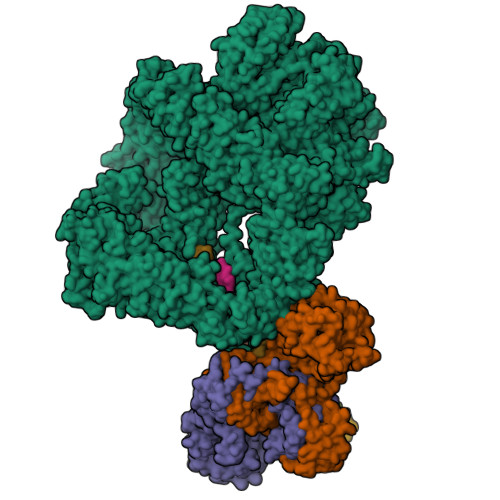
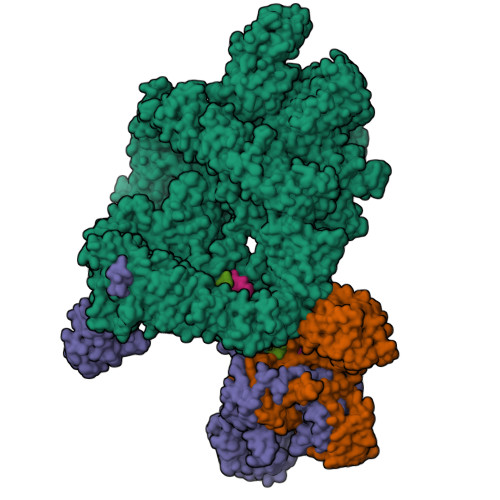
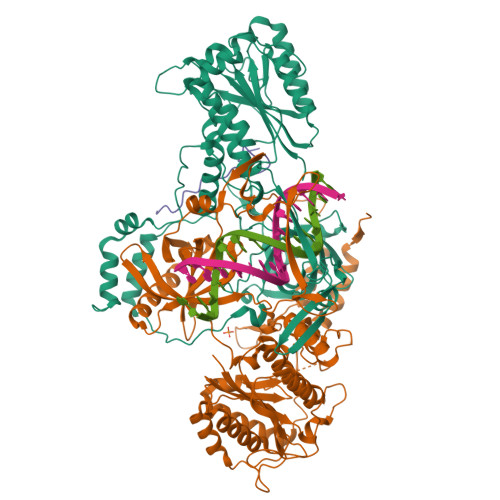
Complex of XLF and heterodimer Ku bound to DNANemoz, C., Legrand, P., Ropars, V., Charbonnier, J.B. (2018) Nat Struct Mol Biol 25: 971-980
Cryo-EM structure of DNA-PK monomerChaplin, A.K., Hardwick, S.W., Chirgadze, D.Y., Blundell, T.L. (2021) Nat Struct Mol Biol 28: 13-19
Cryo-EM structure of DNA-PK dimerChaplin, A.K., Hardwick, S.W., Chirgadze, D.Y., Blundell, T.L. (2021) Nat Struct Mol Biol 28: 13-19
Ku70/80 complex apo formHnizda, A., Tesina, P., Novak, P., Blundell, T.L. (2021) FEBS J 288: 4382-4393
Cryo-EM structure of activated-form DNA-PK (complex VI)Chen, X., Gellert, M., Yang, W. (2021) Mol Cell 81: 801-810.e3
CryoEM structure of inactivated-form DNA-PK (Complex III)Chen, X., Gellert, M., Yang, W. (2021) Mol Cell 81: 801-810.e3
CryoEM structure of inactivated-form DNA-PK (Complex IV)Chen, X., Gellert, M., Yang, W. (2021) Mol Cell 81: 801-810.e3
CryoEM structure of inactivated-form DNA-PK (Complex V)Chen, X., Gellert, M., Yang, W. (2021) Mol Cell 81: 801-810.e3
NHEJ Short-range synaptic complex(2021) Nature 593: 294-298
NHEJ Long-range synaptic complex(2021) Nature 593: 294-298
Cryo-EM structure of NHEJ super-complex (dimer)Chaplin, A.K., Hardwick, S.W., Kefala Stavridi, A., Chirgadze, D.Y., Blundell, T.L. (2021) Mol Cell 81: 3400-3409
Cryo-EM structure of NHEJ super-complex (monomer)Chaplin, A.K., Hardwick, S.W., Kefala Stavridi, A., Chirgadze, D.Y., Blundell, T.L. (2021) Mol Cell 81: 3400-3409
DNA-PK complex of DNA end processingLiu, L., Li, J., Chen, X., Yang, W., Gellert, M. (2022) Mol Cell 82: 177
CryoEM structure of DNA-PK complex VIIChen, X., Liu, L., Gellert, M., Yang, W. (2022) Mol Cell 82: 177
X-Ray studies of Ku70/80 reveal the binding site for IP6Varela, P.F., Charbonnier, J.B. (2023) Nucleic Acids Res 51: 11732-11747
DNA-PK in the active state(2023) Nat Struct Mol Biol 30: 140-147
DNA-PK in the intermediate state(2023) Nat Struct Mol Biol 30: 140-147
Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphateKefala Stavridi, A., Chaplin, A.K., Blundell, T.L. (2023) Nucleic Acids Res 51: 11732-11747
CryoEM structure of Ku heterodimer bound to DNAHardwick, S.W., Kefala-Stavridi, A., Chirgadze, D.Y., Blundell, T.L., Chaplin, A.K. (2023) Nucleic Acids Res 51: 11732-11747
CryoEM structure of Ku heterodimer bound to DNA and PAXXHardwick, S.W., Kefala-Stavridi, A., Chirgadze, D.Y., Blundell, T.L., Chaplin, A.K. (2023) Sci Adv 9: eadg2834-eadg2834
CryoEM structure of Ku heterodimer bound to DNA, PAXX and XLFHardwick, S.W., Kefala-Stavridi, A., Chirgadze, D.Y., Blundell, T.L., Chaplin, A.K. (2023) Sci Adv 9: eadg2834-eadg2834
Vaccinia C16 protein bound to Ku70/Ku80Rivera-Calzada, A., Arribas-Bosacoma, R., Pearl, L.H., Llorca, O. (2022) Nat Commun 13: 7062-7062
1 to 25 of 32 Structures Page 1 of 2 Sort by |