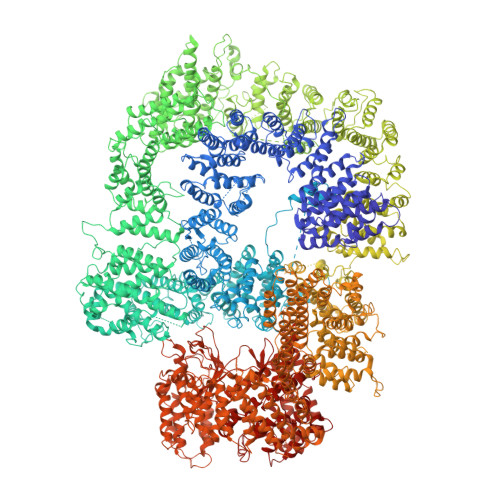
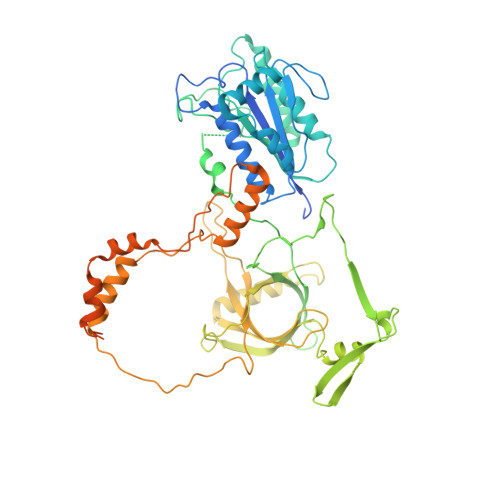
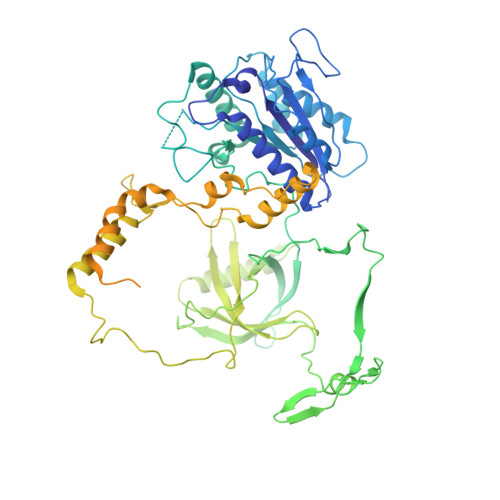
Structure of an activated DNA-PK and its implications for NHEJ.
Chen, X., Xu, X., Chen, Y., Cheung, J.C., Wang, H., Jiang, J., de Val, N., Fox, T., Gellert, M., Yang, W.(2021) Mol Cell 81: 801-810.e3
- PubMed: 33385326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2020.12.015
- Primary Citation Related Structures:
7K0Y, 7K10, 7K11, 7K17, 7K19, 7K1B, 7K1J, 7K1K, 7K1N - PubMed Abstract:
DNA-dependent protein kinase (DNA-PK), like all phosphatidylinositol 3-kinase-related kinases (PIKKs), is composed of conserved FAT and kinase domains (FATKINs) along with solenoid structures made of HEAT repeats. These kinases are activated in response to cellular stress signals, but the mechanisms governing activation and regulation remain unresolved. For DNA-PK, all existing structures represent inactive states with resolution limited to 4.3 Å at best. Here, we report the cryoelectron microscopy (cryo-EM) structures of DNA-PKcs (DNA-PK catalytic subunit) bound to a DNA end or complexed with Ku70/80 and DNA in both inactive and activated forms at resolutions of 3.7 Å overall and 3.2 Å for FATKINs. These structures reveal the sequential transition of DNA-PK from inactive to activated forms. Most notably, activation of the kinase involves previously unknown stretching and twisting within individual solenoid segments and loosens DNA-end binding. This unprecedented structural plasticity of helical repeats may be a general regulatory mechanism of HEAT-repeat proteins.
- Laboratory of Molecular Biology, NIDDK, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: