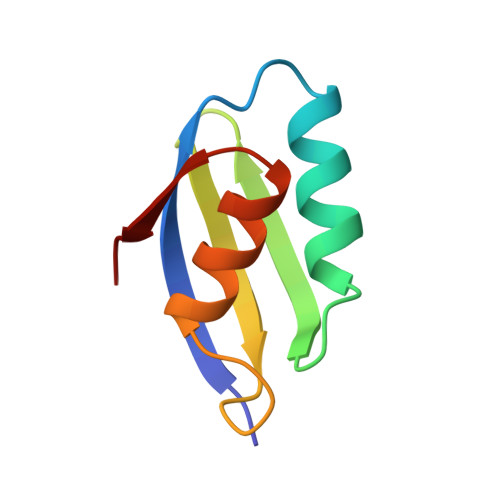
Solution structure of the fourth metal-binding domain from the Menkes copper-transporting ATPase.
Gitschier, J., Moffat, B., Reilly, D., Wood, W.I., Fairbrother, W.J.(1998) Nat Struct Biol 5: 47-54
- PubMed: 9437429 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0198-47
- Primary Citation Related Structures:
1AW0, 2AW0 - PubMed Abstract:
Menkes disease is an X-linked disorder in copper transport that results in death during early childhood. The solution structures of both apo and Ag(I)-bound forms of the fourth metal-binding domain (mbd4) from the Menkes copper-transporting ATPase have been solved. The 72-residue mbd4 has a ferredoxin-like beta alpha beta beta alpha beta fold. Structural differences between the two forms are limited to the metal-binding loop, which is disordered in the apo structure but well ordered in the Ag(I)-bound structure. Ag(I) binds in a linear bicoordinate manner to the two Cys residues of the conserved GMTCxxC motif; Cu(I) likely coordinates in a similar manner. Menkes mbd4 is thus the first bicoordinate copper-binding protein to be characterized structurally. Sequence comparisons with other heavy-metal-binding domains reveal a conserved hydrophobic core and metal-binding motif.
- Howard Hughes Medical Institute, University of California, San Francisco 94143, USA.
Organizational Affiliation: