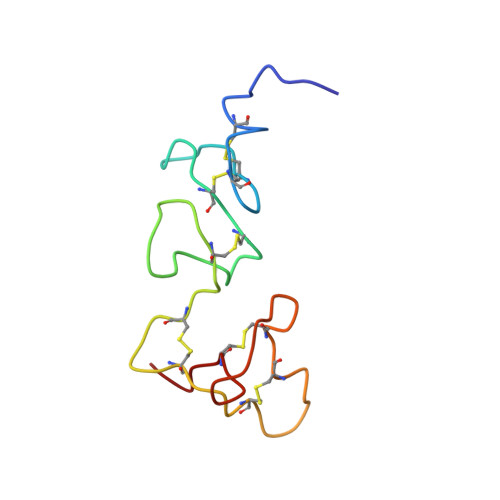
Solution structure of the smallest cofactor-active fragment of thrombomodulin.
Wood, M.J., Sampoli-Benitez, B.A., Komives, E.A.(2000) Nat Struct Biol 7: 200-204
- PubMed: 10700277 Search on PubMed
- DOI: https://doi.org/10.1038/73302
- Primary Citation Related Structures:
1DQB - PubMed Abstract:
A glycosylated fragment of thrombomodulin containing two epidermal growth factor-like domains (TMEGF45) was analyzed by NMR. The 4th-domains structure of this two-domain fragment is similar to that of the individual domain previously determined. The 5th-domain, which has uncrossed disulfide bonds, is not as well determined in the two-domain fragment than the individual domain previously solved. The flexibility of the 5th-domain is consistent with low heteronuclear NOEs. In the individual 5th-domain, Met 388 was disordered, and key thrombin binding residues formed a hydrophobic core. By contrast, in TMEGF45, Met 388 is in the 5th-domain core, positioned by Phe 376 from the 4th-domain. As a result, key thrombin binding residues that were in the core of the individual domain are expelled. Upon thrombin binding, chemical shifts of two residues in the 4th-domain, the three interdomain linker residues, and nearly all of the 5th-domain are perturbed. Thus, TMEGF45 binds thrombin by an induced fit mechanism involving a flexible 5th-domain.
- Department of Chemistry and Biochemistry, University of California, San Diego, 9500 Gilman Dr., La Jolla, California 92093-0359, USA.
Organizational Affiliation: