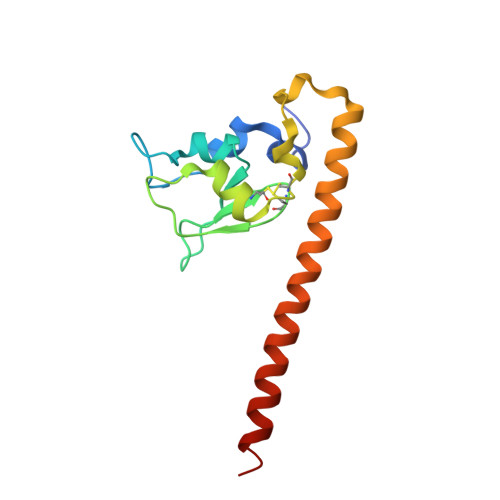
Structure of the human anti-apoptotic protein survivin reveals a dimeric arrangement.
Verdecia, M.A., Huang, H., Dutil, E., Kaiser, D.A., Hunter, T., Noel, J.P.(2000) Nat Struct Biol 7: 602-608
- PubMed: 10876248 Search on PubMed
- DOI: https://doi.org/10.1038/76838
- Primary Citation Related Structures:
1F3H - PubMed Abstract:
Survivin is a 16.5 kDa protein that is expressed during the G2/M phase of the cell cycle and is hypothesized to inhibit a default apoptotic cascade initiated in mitosis. This inhibitory function is coupled to survivin's localization to the mitotic spindle. To begin to address the structural basis of survivin's function, we report the X-ray crystal structure of a recombinant form of full length survivin to 2.58 A resolution. Survivin consists of two defined domains including an N-terminal Zn2+-binding BIR domain linked to a 65 A amphipathic C-terminal alpha-helix. The crystal structure reveals an extensive dimerization interface along a hydrophobic surface on the BIR domain of each survivin monomer. A basic patch acting as a sulfate/phosphate-binding module, an acidic cluster projecting off the BIR domain, and a solvent-accessible hydrophobic surface residing on the C-terminal amphipathic helix, are suggestive of functional protein-protein interaction surfaces.
- Structural Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, California 92037, USA.
Organizational Affiliation: