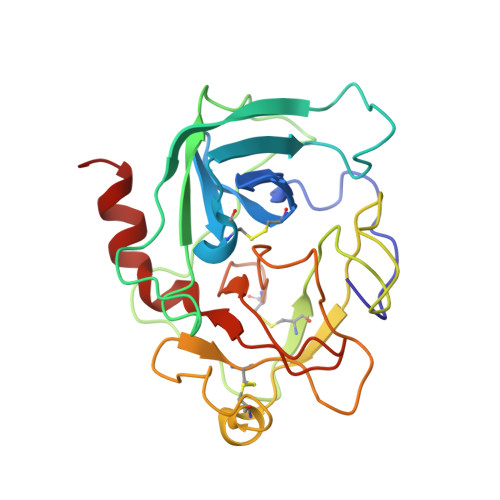
Crystal structure of the caspase activator human granzyme B, a proteinase highly specific for an Asp-P1 residue.
Estebanez-Perpina, E., Fuentes-Prior, P., Belorgey, D., Braun, M., Kiefersauer, R., Maskos, K., Huber, R., Rubin, H., Bode, W.(2000) Biol Chem 381: 1203-1214
- PubMed: 11209755 Search on PubMed
- DOI: https://doi.org/10.1515/BC.2000.148
- Primary Citation Related Structures:
1FQ3 - PubMed Abstract:
Granzyme B is the prototypic member of the granzymes, a family of trypsin-like serine proteinases localized in the dense cytoplasmic granules of activated natural killer cells and cytotoxic T lymphocytes. Granzyme B directly triggers apoptosis in target cells by activating the caspase pathway, and has been implicated in the etiology of rheumatoid arthritis. Human granzyme B expressed in a baculovirus system has been crystallized without inhibitor and its structure has been determined to 3.1 A resolution, after considerably improving the diffraction power of the crystals by controlled humidity changes. The granzyme B structure reveals an overall fold similar to that found in cathepsin G and human chymase. The guanidinium group of Arg226, anchored at the back of the S1-specificity pocket, can form a salt bridge with the P1-Asp side chain of a bound peptide substrate. The architecture of the substrate binding site of granzyme B appears to be designed to accommodate and cleave hexapeptides such as the sequence Ile-Glu-Thr-Asp-/Ser-Gly present in the activation site of pro-caspase-3, a proven physiological substrate of granzyme B. These granzyme B crystals, with fully accessible active sites, are well suited for soaking with small synthetic inhibitors that might be used for a treatment of chronic inflammatory disorders.
- Abteilung für Strukturforschung, Max-Planck-Institut für Biochemie, Planegg-Martinsried, Germany.
Organizational Affiliation: