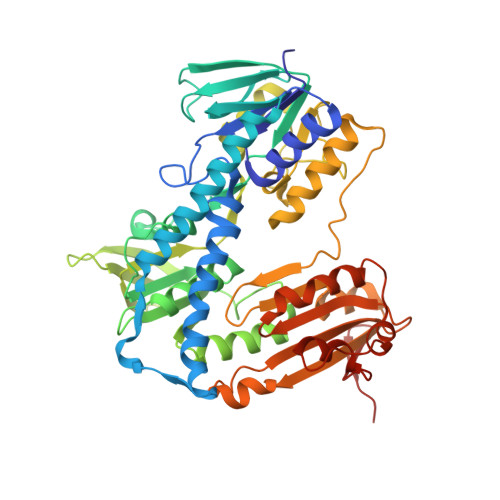
Two Interacting Binding Sites for Quinacrine Derivatives in the Active Site of Trypanothione Reductase: A Template for Drug Design
Saravanamuthu, A., Vickers, T.J., Bond, C.S., Peterson, M.R., Hunter, W.N., Fairlamb, A.H.(2004) J Biological Chem 279: 29493
- PubMed: 15102853
- DOI: https://doi.org/10.1074/jbc.M403187200
- Primary Citation of Related Structures:
1GXF - PubMed Abstract:
Trypanothione reductase is a key enzyme in the trypanothione-based redox metabolism of pathogenic trypanosomes. Because this system is absent in humans, being replaced with glutathione and glutathione reductase, it offers a target for selective inhibition. The rational design of potent inhibitors requires accurate structures of enzyme-inhibitor complexes, but this is lacking for trypanothione reductase. We therefore used quinacrine mustard, an alkylating derivative of the competitive inhibitor quinacrine, to probe the active site of this dimeric flavoprotein. Quinacrine mustard irreversibly inactivates Trypanosoma cruzi trypanothione reductase, but not human glutathione reductase, in a time-dependent manner with a stoichiometry of two inhibitors bound per monomer. The rate of inactivation is dependent upon the oxidation state of trypanothione reductase, with the NADPH-reduced form being inactivated significantly faster than the oxidized form. Inactivation is slowed by clomipramine and a melarsen oxide-trypanothione adduct (both are competitive inhibitors) but accelerated by quinacrine. The structure of the trypanothione reductase-quinacrine mustard adduct was determined to 2.7 A, revealing two molecules of inhibitor bound in the trypanothione-binding site. The acridine moieties interact with each other through pi-stacking effects, and one acridine interacts in a similar fashion with a tryptophan residue. These interactions provide a molecular explanation for the differing effects of clomipramine and quinacrine on inactivation by quinacrine mustard. Synergism with quinacrine occurs as a result of these planar acridines being able to stack together in the active site cleft, thereby gaining an increased number of binding interactions, whereas antagonism occurs with nonplanar molecules, such as clomipramine, where stacking is not possible.
- Division of Biological Chemistry and Molecular Microbiology, The Wellcome Trust Biocentre, School of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: