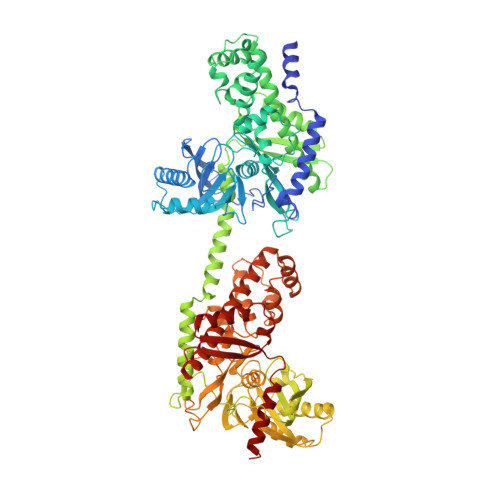
The mechanism of regulation of hexokinase: new insights from the crystal structure of recombinant human brain hexokinase complexed with glucose and glucose-6-phosphate.
Aleshin, A.E., Zeng, C., Bourenkov, G.P., Bartunik, H.D., Fromm, H.J., Honzatko, R.B.(1998) Structure 6: 39-50
- PubMed: 9493266 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00006-9
- Primary Citation Related Structures:
1HKB - PubMed Abstract:
Hexokinase I is the pacemaker of glycolysis in brain tissue. The type I isozyme exhibits unique regulatory properties in that physiological levels of phosphate relieve potent inhibition by the product, glucose-6-phosphate (Gluc-6-P). The 100 kDa polypeptide chain of hexokinase I consists of a C-terminal (catalytic) domain and an N-terminal (regulatory) domain. Structures of ligated hexokinase I should provide a basis for understanding mechanisms of catalysis and regulation at an atomic level. The complex of human hexokinase I with glucose and Gluc-6-P (determined to 2.8 A resolution) is a dimer with twofold molecular symmetry. The N- and C-terminal domains of one monomer interact with the C- and N-terminal domains, respectively, of the symmetry-related monomer. The two domains of a monomer are connected by a single alpha helix and each have the fold of yeast hexokinase. Salt links between a possible cation-binding loop of the N-terminal domain and a loop of the C-terminal domain may be important to regulation. Each domain binds single glucose and Gluc-6-P molecules in proximity to each other. The 6-phosphoryl group of bound Gluc-6-P at the C-terminal domain occupies the putative binding site for ATP, whereas the 6-phosphoryl group at the N-terminal domain may overlap the binding site for phosphate. The binding synergism of glucose and Gluc-6-P probably arises out of the mutual stabilization of a common (glucose-bound) conformation of hexokinase I. Conformational changes in the N-terminal domain in response to glucose, phosphate, and/or Gluc-6-P may influence the binding of ATP to the C-terminal domain.
- Department of Biochemistry and Biophysics, Iowa State University, Ames 50011, USA.
Organizational Affiliation: