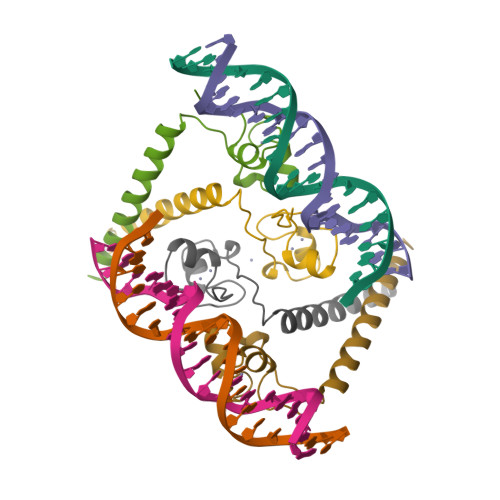
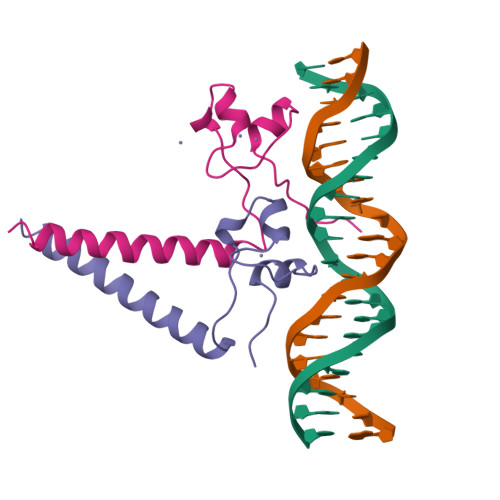
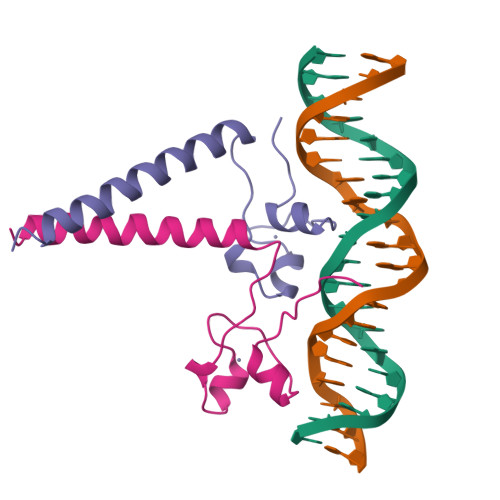
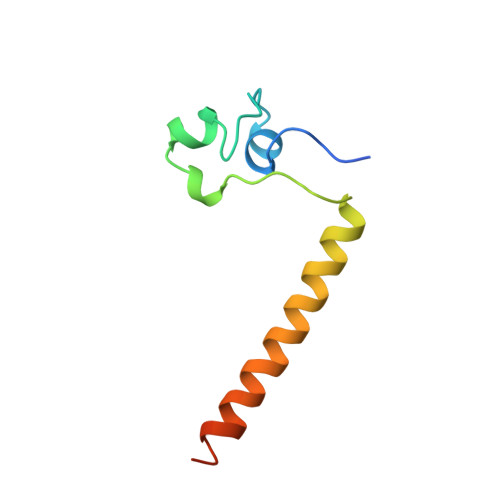
Structure of a HAP1-DNA complex reveals dramatically asymmetric DNA binding by a homodimeric protein.
King, D.A., Zhang, L., Guarente, L., Marmorstein, R.(1999) Nat Struct Biol 6: 64-71
- PubMed: 9886294
- DOI: https://doi.org/10.1038/4940
- Primary Citation of Related Structures:
1HWT - PubMed Abstract:
HAP1 is a member of a family of fungal transcription factors that contain a Zn2Cys6 binuclear cluster domain and bind as homodimers to sequences containing two DNA half sites. We have determined the 2.5 A crystal structure of HAP1 bound to a cognate upstream activation sequence from the CYC7 gene. The structure reveals that HAP1 is bound in a dramatically asymmetric manner to the DNA target. This asymmetry aligns the Zn2Cys6 domains in a tandem head-to-tail fashion to contact two DNA half sites, positions an N-terminal arm of one of the protein subunits to interact with the inter-half site base pairs in the DNA minor groove, and suggests a mechanism by which DNA-binding facilitates asymmetric dimerization by HAP1. Comparisons with the DNA complexes of the related GAL4, PPR1 and PUT3 proteins illustrate how a conserved protein domain can be reoriented to recognize DNA half sites of different polarities and how homodimeric proteins adopt dramatically asymmetric structures to recognize cognate DNA targets.
Organizational Affiliation:
The Wistar Institute and The Department of Chemistry, University of Pennsylvania, Philadelphia 19104, USA.