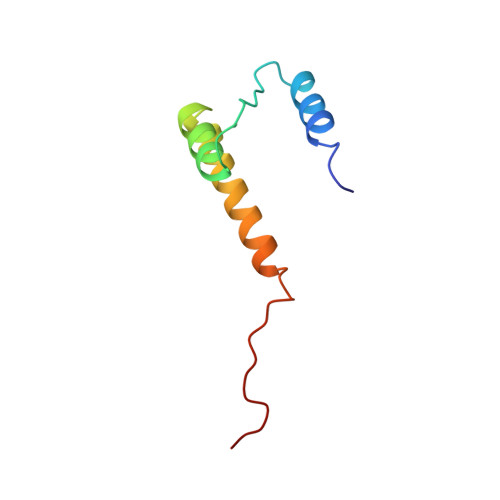
Structure of a trimeric domain of the MHC class II-associated chaperonin and targeting protein Ii.
Jasanoff, A., Wagner, G., Wiley, D.C.(1998) EMBO J 17: 6812-6818
- PubMed: 9843486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/17.23.6812
- Primary Citation Related Structures:
1IIE - PubMed Abstract:
The invariant chain (Ii) plays a critical role in MHC class II antigen processing by stabilizing peptide-free class II alphabeta heterodimers in a nonameric (alphabetaIi)3 complex soon after their synthesis and directing transport of the complex from the endoplasmic reticulum to compartments where peptide loading of class II takes place. Loading progresses following Ii proteolysis and via an intermediate complex of MHC class II with an Ii-derived peptide, CLIP. CLIP is substituted by exogenous peptidic fragments in an exchange reaction catalyzed by HLA-DM. The CLIP region of Ii, roughly residues 81-104, is one of two segments shown to interact with class II HLA-DR molecules. The other segment, Ii 118-216, is C-terminal to CLIP, mediates trimerization of the ectodomain of Ii and interferes with DM/class II binding. Here we report the three-dimensional structure of this trimeric domain, determined by nuclear magnetic resonance (NMR) studies of a 27 kDa trimer of human Ii 118-192. The cylindrical shape of the molecule and the mapping of conserved residues delimit surfaces which may be important for interactions between Ii and class II molecules.
- Department of Molecular and Cellular Biology, Harvard Medical School, Cambridge, MA 02138, USA. jasanoff@crystal.harvard.edu
Organizational Affiliation: