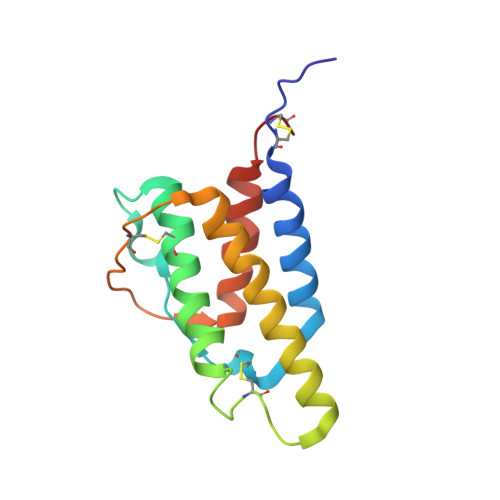
The high-resolution, three-dimensional solution structure of human interleukin-4 determined by multidimensional heteronuclear magnetic resonance spectroscopy.
Powers, R., Garrett, D.S., March, C.J., Frieden, E.A., Gronenborn, A.M., Clore, G.M.(1993) Biochemistry 32: 6744-6762
- PubMed: 8329398 Search on PubMed
- DOI: https://doi.org/10.1021/bi00077a030
- Primary Citation Related Structures:
1ITI - PubMed Abstract:
The high-resolution three-dimensional solution structure of recombinant human interleukin-4 (IL-4), a protein of approximately 15 kDa which plays a key role in the regulation of B and T lymphocytes, has been determined using three- and four-dimensional heteronuclear NMR spectroscopy. The structure is based on a total of 2973 experimental NMR restraints, comprising 2515 approximate interproton distance restraints, 102 distance restraints for 51 backbone hydrogen bonds, and 356 torsion angle restraints. A total of 30 structures was calculated by means of hybrid distance geometry-simulated annealing, and the atomic rms distribution about the mean coordinate positions for residues 8-129 is 0.44 +/- 0.03 A for the backbone atoms, 0.83 +/- 0.03 A for all atoms, and 0.51 +/- 0.04 A for all atoms excluding disordered side chains. The N- and C-terminal residues (1-7 and 130-133, respectively) appear to be disordered. The structure of IL-4 is dominated by a left-handed four-helix bundle with an unusual topology comprising two overhand connections. The linker elements between the helices are formed by either long loops, small helical turns, or short strands. The latter include a mini anti-parallel beta-sheet. A best fit superposition of the NMR structure of IL-4 with the 2.25 A resolution crystal structure [Wlodawer, A., Pavlovsky, A., & Gutschina, A. (1992) FEBS Lett. 309, 59-64] yields a backbone atomic rms difference of 1.37 A which can be mainly attributed to tighter packing of the helices in the crystal structure. This is indicated by an approximately 20% reduction in the axial separation of three pairs of helices (alpha A-alpha C, alpha A-alpha D, and alpha C-alpha D) in the crystal structure relative to the NMR structure and may reflect the greater flexibility of the molecule in solution which is reduced in the crystal due to intermolecular contacts. Comparison of the NMR structure of IL-4 with the X-ray structures of two other related proteins, granulocyte-macrophage colony stimulating factor [Diedrichs, K., Boone, T., & Karplus, P. A. (1992) Science 254, 1779-1782] and human growth hormone [de Vos, A. M., Ultsch, M., & Kossiakoff, A. A. (1992) Science 255, 306-312], that bind to the same hematopoietic superfamily of cell surface receptors reveals a remarkably similar topological fold, despite the absence of any significant overall sequence identity, and substantial differences in the relative lengths of the helices, the lengths and the nature of the various connecting elements, and the pattern and number of disulfide bridges.(ABSTRACT TRUNCATED AT 400 WORDS)
- Laboratory of Chemical Physics, National Institutes of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892.
Organizational Affiliation: