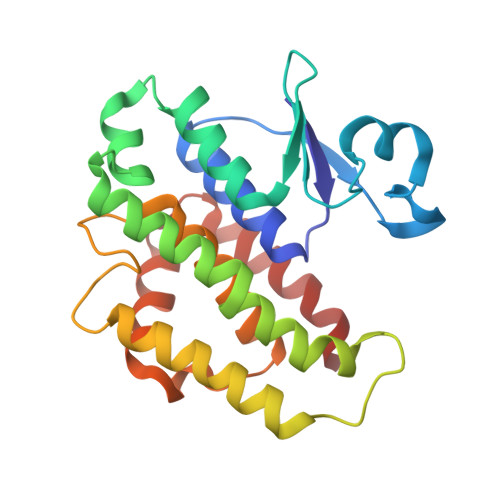
The crystal structures of glutathione S-transferases isozymes 1-3 and 1-4 from Anopheles dirus species B.
Oakley, A.J., Harnnoi, T., Udomsinprasert, R., Jirajaroenrat, K., Ketterman, A.J., Wilce, M.C.(2001) Protein Sci 10: 2176-2185
- PubMed: 11604524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.ps.21201
- Primary Citation Related Structures:
1JLV, 1JLW - PubMed Abstract:
Glutathione S-transferases (GSTs) are dimeric proteins that play an important role in cellular detoxification. Four GSTs from the mosquito Anopheles dirus species B (Ad), an important malaria vector in South East Asia, are produced by alternate splicing of a single transcription product and were previously shown to have detoxifying activity towards pesticides such as DDT. We have determined the crystal structures for two of these alternatively spliced proteins, AdGST1-3 (complexed with glutathione) and AdGST1-4 (apo form), at 1.75 and 2.45 A resolution, respectively. These GST isozymes show differences from the related GST from the Australian sheep blowfly Lucilia cuprina; in particular, the presence of a C-terminal helix forming part of the active site. This helix causes the active site of the Anopheles GSTs to be enclosed. The glutathione-binding helix alpha2 and flanking residues are disordered in the AdGST1-4 (apo) structure, yet ordered in the AdGST1-3 (GSH-bound) structure, suggesting that insect GSTs operate with an induced fit mechanism similar to that found in the plant phi- and human pi-class GSTs. Despite the high overall sequence identities, the active site residues of AdGST1-4 and AdGST1-3 have different conformations.
- Department of Pharmacology/Crystallography Centre, University of Western Australia, Crawley 6009, Australia.
Organizational Affiliation: