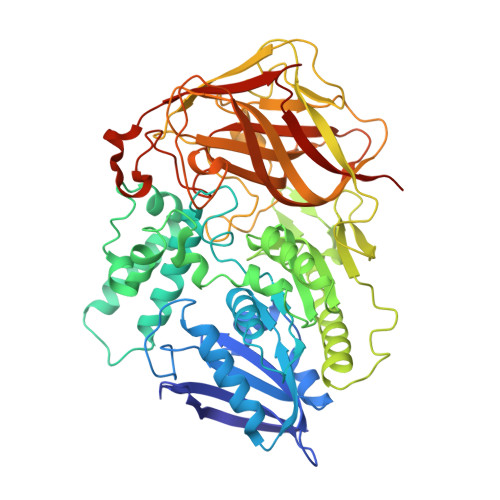
Crystal structure of a bacterial cocaine esterase.
Larsen, N.A., Turner, J.M., Stevens, J., Rosser, S.J., Basran, A., Lerner, R.A., Bruce, N.C., Wilson, I.A.(2002) Nat Struct Biol 9: 17-21
- PubMed: 11742345 Search on PubMed
- DOI: https://doi.org/10.1038/nsb742
- Primary Citation Related Structures:
1JU3, 1JU4 - PubMed Abstract:
Here we report the first structure of a cocaine-degrading enzyme. The bacterial esterase, cocE, hydrolyzes pharmacologically active (-)-cocaine to a non-psychoactive metabolite with a rate faster than any other reported cocaine esterase (kcat = 7.8 s-1 and KM = 640 nM). Because of the high catalytic proficiency of cocE, it is an attractive candidate for novel protein-based therapies for cocaine overdose. The crystal structure of cocE, solved by multiple anomalous dispersion (MAD) methods, reveals that cocE is a serine esterase composed of three domains: (i) a canonical alpha/beta hydrolase fold (ii) an alpha-helical domain that caps the active site and (iii) a jelly-roll-like beta-domain that interacts extensively with the other two domains. The active site was identified within the interface of all three domains by analysis of the crystal structures of transition state analog adduct and product complexes, which were refined at 1.58 A and 1.63 A resolution, respectively. These structural studies suggest that substrate recognition arises partly from interactions between the benzoyl moiety of cocaine and a highly evolved specificity pocket.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Rd., La Jolla, California 92037, USA.
Organizational Affiliation: