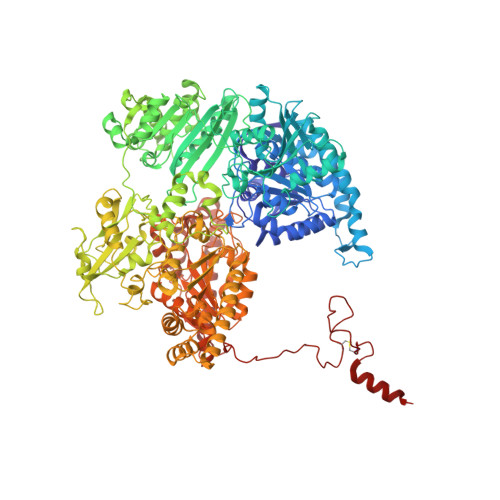
Crystal structure of the free radical intermediate of pyruvate:ferredoxin oxidoreductase.
Chabriere, E., Vernede, X., Guigliarelli, B., Charon, M.H., Hatchikian, E.C., Fontecilla-Camps, J.C.(2001) Science 294: 2559-2563
- PubMed: 11752578 Search on PubMed
- DOI: https://doi.org/10.1126/science.1066198
- Primary Citation Related Structures:
1KEK - PubMed Abstract:
In anaerobic organisms, the decarboxylation of pyruvate, a crucial component of intermediary metabolism, is catalyzed by the metalloenzyme pyruvate: ferredoxin oxidoreductase (PFOR) resulting in the generation of low potential electrons and the subsequent acetylation of coenzyme A (CoA). PFOR is the only enzyme for which a stable acetyl thiamine diphosphate (ThDP)-based free radical reaction intermediate has been identified. The 1.87 A-resolution structure of the radical form of PFOR from Desulfovibrio africanus shows that, despite currently accepted ideas, the thiazole ring of the ThDP cofactor is markedly bent, indicating a drastic reduction of its aromaticity. In addition, the bond connecting the acetyl group to ThDP is unusually long, probably of the one-electron type already described for several cation radicals but not yet found in a biological system. Taken together, our data, along with evidence from the literature, suggest that acetyl-CoA synthesis by PFOR proceeds via a condensation mechanism involving acetyl (PFOR-based) and thiyl (CoA-based) radicals.
- Laboratoire de Cristallographie et Cristallogenèse des Protéines, Institut de Biologie Structurale Jean-Pierre Ebel, Commissariat à l'Energie Atomique, Université Joseph Fourier, CNRS, 41, rue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: