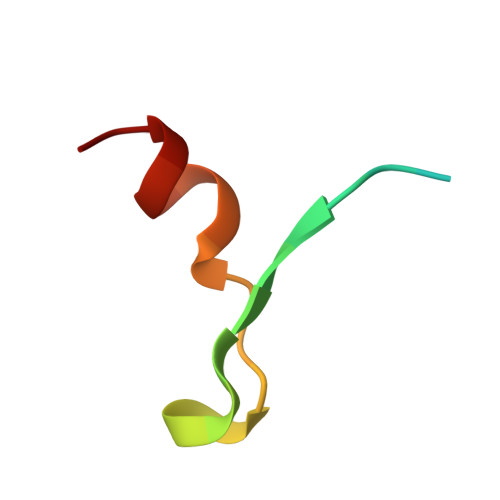
Recognition of eIF4G by rotavirus NSP3 reveals a basis for mRNA circularization.
Groft, C.M., Burley, S.K.(2002) Mol Cell 9: 1273-1283
- PubMed: 12086624 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(02)00555-5
- Primary Citation Related Structures:
1LJ2 - PubMed Abstract:
Rotaviruses, segmented double-stranded RNA viruses, co-opt the eukaryotic translation machinery with the aid of nonstructural protein 3 (NSP3), a rotaviral functional homolog of the cellular poly(A) binding protein (PABP). NSP3 binds to viral mRNA 3' consensus sequences and circularizes mRNA via interactions with eIF4G. Here, we present the X-ray structure of the C-terminal domain of NSP3 (NSP3-C) recognizing a fragment of eIF4GI. Homodimerization of NSP3-C yields a symmetric, elongated, largely alpha-helical structure with two hydrophobic eIF4G binding pockets at the dimer interface. Site-directed mutagenesis and isothermal titration calorimetry documented that NSP3 and PABP use analogous eIF4G recognition strategies, despite marked differences in tertiary structure.
- Laboratory of Molecular Biophysics, New York, NY 10021, USA.
Organizational Affiliation: