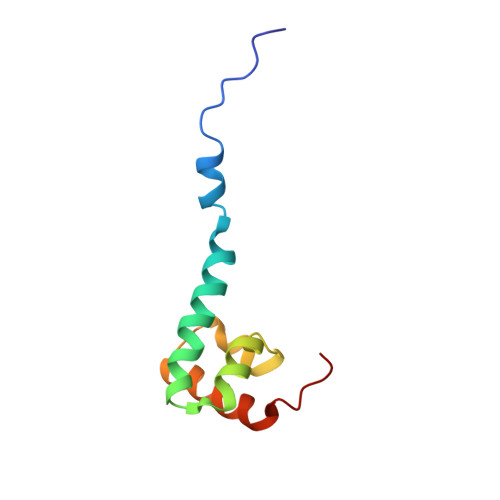
Solution Structure of the C-terminal Domain from poly(A)-binding protein in Trypanosoma cruzi: A vegetal PABC domain
Siddiqui, N., Kozlov, G., D'Orso, I., Trempe, J.F., Gehring, K.(2003) Protein Sci 12: 1925-1933
- PubMed: 12930992
- DOI: https://doi.org/10.1110/ps.0390103
- Primary Citation Related Structures:
1NMR - PubMed Abstract:
PABC is a phylogenetically conserved peptide-binding domain primarily found within the C terminus of poly(A)-binding proteins (PABPs). This domain recruits a series of translation factors including poly(A)-interacting proteins (Paip1 and Paip2) and release factor 3 (RF3/GSPT) to the initiation complex on mRNA. Here, we determine the solution structure of the Trypanosoma cruzi PABC domain (TcPABC), a representative of the vegetal class of PABP proteins. TcPABC is similar to human PABC (hPABC) and consists of five alpha-helices, in contrast to the four helices observed in PABC domains from yeast (yPABC) and hyper plastic disk proteins (hHYD). A mobile N-terminal helix is observed in TcPABC that does not pack against the core of the protein, as found in hPABC. Characteristic to all PABC domains, the last four helices of TcPABC fold into a right-handed super coil. TcPABC demonstrates high-affinity binding to PABP interacting motif-2 (PAM-2) and reveals a peptide-binding surface homologous to that of hPABC. Our results demonstrate the last four helices in TcPABC are sufficient for peptide recognition and we predict a similar binding mode in PABC domains. Furthermore, these results point to the presence of putative PAM-2 site-containing proteins in trypanosomes.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6, Canada.
Organizational Affiliation: