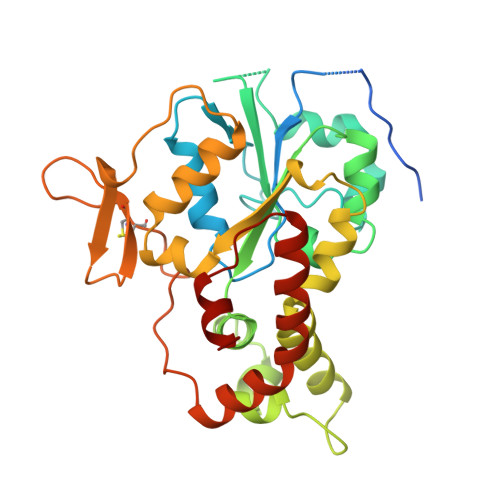
Crystal structure of the sulfotransferase domain of human heparan sulfate N-deacetylase/ N-sulfotransferase 1.
Kakuta, Y., Sueyoshi, T., Negishi, M., Pedersen, L.C.(1999) J Biological Chem 274: 10673-10676
- PubMed: 10196134 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.16.10673
- Primary Citation Related Structures:
1NST - PubMed Abstract:
Heparan sulfate N-deacetylase/N-sulfotransferase (HSNST) catalyzes the first and obligatory step in the biosynthesis of heparan sulfates and heparin. The crystal structure of the sulfotransferase domain (NST1) of human HSNST-1 has been determined at 2.3-A resolution in a binary complex with 3'-phosphoadenosine 5'-phosphate (PAP). NST1 is approximately spherical with an open cleft, and consists of a single alpha/beta fold with a central five-stranded parallel beta-sheet and a three-stranded anti-parallel beta-sheet bearing an interstrand disulfide bond. The structural regions alpha1, alpha6, beta1, beta7, 5'-phosphosulfate binding loop (between beta1 and alpha1), and a random coil (between beta8 and alpha13) constitute the PAP binding site of NST1. The alpha6 and random coil (between beta2 and alpha2), which form an open cleft near the 5'-phosphate of the PAP molecule, may provide interactions for substrate binding. The conserved residue Lys-614 is in position to form a hydrogen bond with the bridge oxygen of the 5'-phosphate.
- Pharmacogenetics Section, Laboratory of Reproductive and Developmental Toxicology, NIEHS, National Institutes of Health, Research Triangle Park, North Carolina 27709, USA.
Organizational Affiliation: