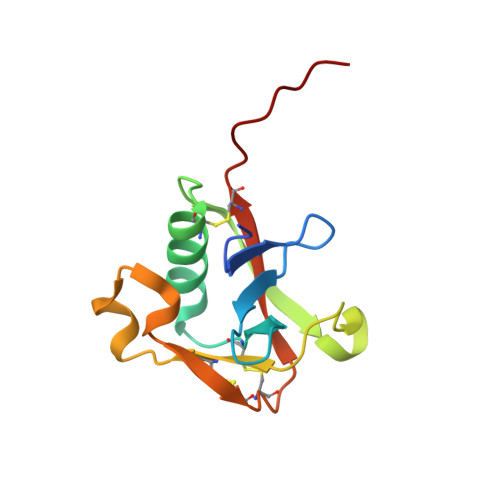
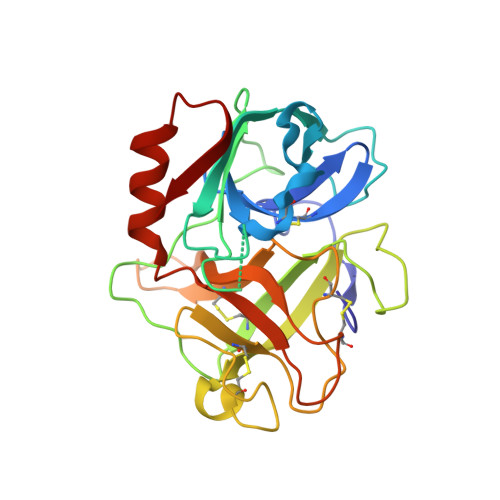
Dissecting and designing inhibitor selectivity determinants at the S1 site using an artificial Ala190 protease (Ala190 uPA).
Katz, B.A., Luong, C., Ho, J.D., Somoza, J.R., Gjerstad, E., Tang, J., Williams, S.R., Verner, E., Mackman, R.L., Young, W.B., Sprengeler, P.A., Chan, H., Mortara, K., Janc, J.W., McGrath, M.E.(2004) J Mol Biology 344: 527-547
- PubMed: 15522303 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.09.032
- Primary Citation Related Structures:
1O5A, 1O5B, 1O5C, 1O5D, 1O5E, 1O5F, 1O5G - PubMed Abstract:
A site-directed mutant of the serine protease urokinase-type plasminogen activator (uPA), was produced to assess the contribution of the Ser190 side-chain to the affinity and selectivity of lead uPA inhibitors in the absence of other differences present in comparisons of natural proteases. Crystallography and enzymology involving WT and Ala190 uPA were used to calculate free energy binding contributions of hydrogen bonds involving the Ser190 hydroxyl group (O(gamma)(Ser190)) responsible for the remarkable selectivity of 6-halo-5-amidinoindole and 6-halo-5-amidinobenzimidazole inhibitors toward uPA and against natural Ala190 protease anti-targets. Crystal structures of uPA complexes of novel, active site-directed arylguanidine and 2-aminobenzimidazole inhibitors of WT uPA, together with associated K(i) values for WT and Ala190 uPA, also indicate a significant role of Ser190 in the binding of these classes of uPA inhibitors. Structures and associated K(i) values for a lead inhibitor (CA-11) bound to uPA and to five other proteases, as well as for other leads bound to multiple proteases, help reveal the features responsible for the potency (K(i)=11nM) and selectivity of the remarkably small inhibitor, CA-11. The 6-fluoro-5-amidinobenzimidzole, CA-11, is more than 1000-fold selective against natural Ala190 protease anti-targets, and more than 100-fold selective against other Ser190 anti-targets.
- Celera, 180 Kimball Way, South San Francisco, CA 94080, USA. brad.katz@celera.com
Organizational Affiliation: