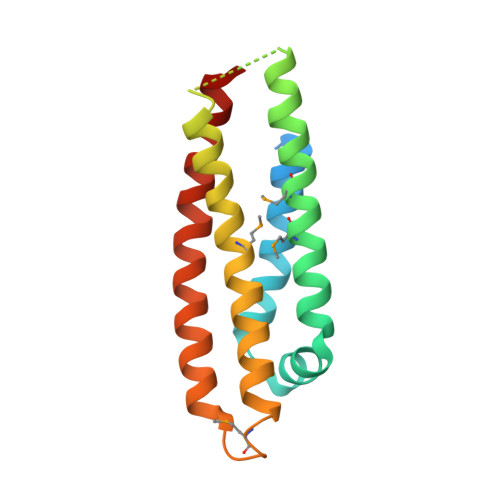
Conformational flexibility in the apolipoprotein E amino-terminal domain structure determined from three new crystal forms: implications for lipid binding.
Segelke, B.W., Forstner, M., Knapp, M., Trakhanov, S.D., Parkin, S., Newhouse, Y.M., Bellamy, H.D., Weisgraber, K.H., Rupp, B.(2000) Protein Sci 9: 886-897
- PubMed: 10850798 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.9.5.886
- Primary Citation Related Structures:
1BZ4, 1OR2, 1OR3 - PubMed Abstract:
An amino-terminal fragment of human apolipoprotein E3 (residues 1-165) has been expressed and crystallized in three different crystal forms under similar crystallization conditions. One crystal form has nearly identical cell dimensions to the previously reported orthorhombic (P2(1)2(1)2(1)) crystal form of the amino-terminal 22 kDa fragment of apolipoprotein E (residues 1-191). A second orthorhombic crystal form (P2(1)2(1)2(1) with cell dimensions differing from the first form) and a trigonal (P3(1)21) crystal form were also characterized. The structures of the first orthorhombic and the trigonal form were determined by seleno-methionine multiwavelength anomalous dispersion, and the structure of the second orthorhombic form was determined by molecular replacement using the structure from the trigonal form as a search model. A combination of modern experimental and computational techniques provided high-quality electron-density maps, which revealed new features of the apolipoprotein E structure, including an unambiguously traced loop connecting helices 2 and 3 in the four-helix bundle and a number of multiconformation side chains. The three crystal forms contain a common intermolecular, antiparallel packing arrangement. The electrostatic complimentarity observed in this antiparallel packing resembles the interaction of apolipoprotein E with the monoclonal antibody 2E8 and the low density lipoprotein receptor. Superposition of the model structures from all three crystal forms reveals flexibility and pronounced kinks in helices near one end of the four-helix bundle. This mobility at one end of the molecule provides new insights into the structural changes in apolipoprotein E that occur with lipid association.
- Lawrence Livermore National Laboratory, Biology and Biotechnology Research Program, University of California, Livermore 94550, USA.
Organizational Affiliation: