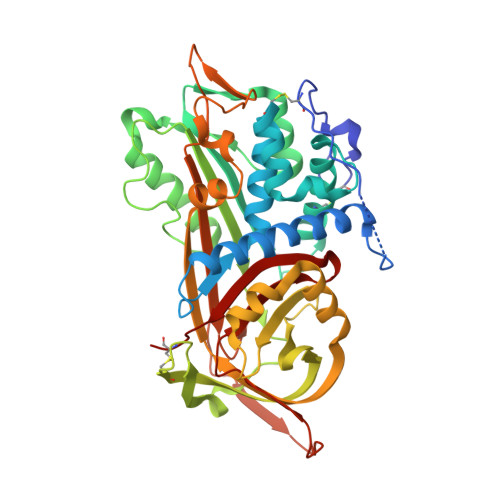
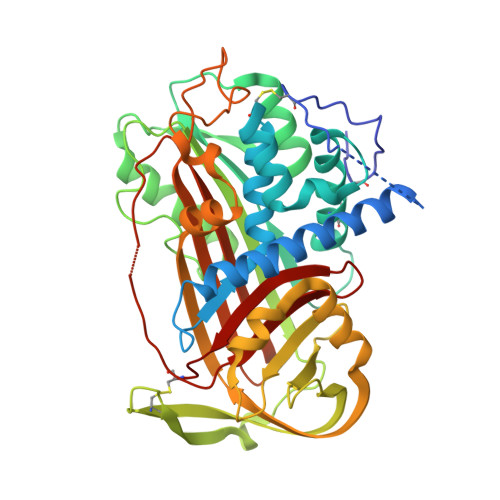
The influence of hinge region residue Glu-381 on antithrombin allostery and metastability
Johnson, D.J.D., Huntington, J.A.(2004) J Biological Chem 279: 4913-4921
- PubMed: 14623882 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M311644200
- Primary Citation Related Structures:
1OYH - PubMed Abstract:
Antithrombin becomes an efficient inhibitor of factor Xa and thrombin by binding a specific pentasaccharide sequence found on a small fraction of the heparan sulfate proteoglycans lining the microvaculature. In the structure of native antithrombin, the reactive center loop is restrained due to the insertion of its hinge region into the main beta-sheet A, whereas in the heparin-activated state the reactive center loop is freed from beta-sheet A. In both structures, hinge region residue Glu-381 makes several stabilizing contacts. To determine the role of these contacts in the allosteric mechanism of antithrombin activation, we replaced Glu-381 with an alanine. This variant is less active toward its target proteases than control antithrombin, due to a perturbation of the equilibrium between the two forms, and to an increase in stoichiometry of inhibition. Pentasaccharide binding affinity is reduced 4-fold due to an increase in the off-rate. These data suggest that the main role of Glu-381 is to stabilize the activated conformation. Stability studies also showed that the E381A variant is resistant to continued insertion of its reactive center loop upon incubation at 50 degrees C, suggesting new stabilizing interactions in the native structure. To test this hypothesis, and to aid in the interpretation of the kinetic data we solved to 2.6 A the structure of the variant. We conclude that wild-type Glu-381 interactions stabilize the activated state and decreases the energy barrier to full loop insertion.
- University of Cambridge, Department of Haematology, Cambridge Institute for Medical Research, Division of Structural Medicine, Thrombosis Research Unit, Wellcome Trust MRC Building, Hills Road, Cambridge CB2 2XY, United Kingdom.
Organizational Affiliation: