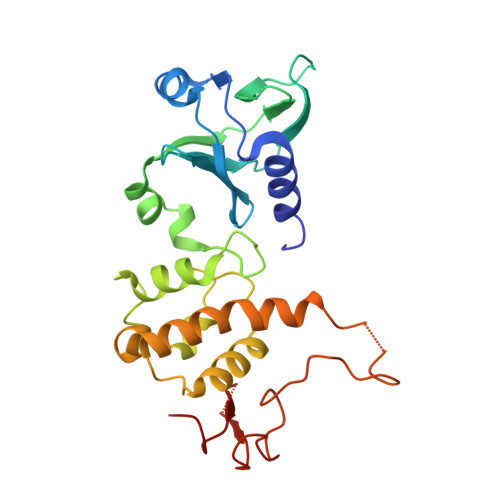
Structure of the uncomplexed DNA repair enzyme endonuclease VIII indicates significant interdomain flexibility.
Golan, G., Zharkov, D.O., Feinberg, H., Fernandes, A.S., Zaika, E.I., Kycia, J.H., Grollman, A.P., Shoham, G.(2005) Nucleic Acids Res 33: 5006-5016
- PubMed: 16145054
- DOI: https://doi.org/10.1093/nar/gki796
- Primary Citation of Related Structures:
1Q39, 1Q3B, 1Q3C - PubMed Abstract:
Escherichia coli endonuclease VIII (Nei) excises oxidized pyrimidines from DNA. It shares significant sequence homology and similar mechanism with Fpg, a bacterial 8-oxoguanine glycosylase. The structure of a covalent Nei-DNA complex has been recently determined, revealing critical amino acid residues which are important for DNA binding and catalysis. Several Fpg structures have also been reported; however, analysis of structural dynamics of Fpg/Nei family proteins has been hindered by the lack of structures of uncomplexed and DNA-bound enzymes from the same source. We report a 2.8 A resolution structure of free wild-type Nei and two structures of its inactive mutants, Nei-E2A (2.3 A) and Nei-R252A (2.05 A). All three structures are virtually identical, demonstrating that the mutations did not affect the overall conformation of the protein in its free state. The structures show a significant conformational change compared with the Nei structure in its complex with DNA, reflecting a approximately 50 degrees rotation of the two main domains of the enzyme. Such interdomain flexibility has not been reported previously for any DNA glycosylase and may present the first evidence for a global DNA-induced conformational change in this class of enzymes. Several local but functionally relevant structural changes are also evident in other parts of the enzyme.
- Department of Inorganic Chemistry and the Laboratory for Structural Chemistry and Biology, The Hebrew University of Jerusalem Jerusalem 91904, Israel.
Organizational Affiliation: