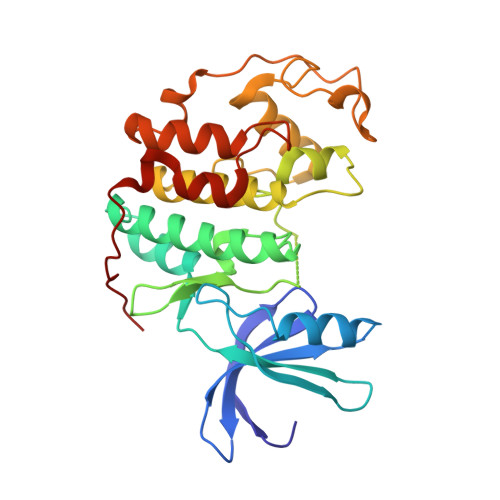
A new series of potent oxindole inhibitors of CDK2
Luk, K.-C., Simcox, M.E., Schutt, A., Rowan, K., Thompson, T., Chen, Y., Kammlott, U., DePinto, W., Dunten, P., Dermatakis, A.(2004) Bioorg Med Chem Lett 14: 913-917
- PubMed: 15012993 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2003.12.009
- Primary Citation Related Structures:
1R78 - PubMed Abstract:
A novel series of oxindole-type inhibitors of CDK2 that have heteroatom substituted alkynyl moieties at their C-4 position is described. These novel 4-alkynyl-substituted inhibitors have superior potency relative to their parent compound in free enzyme and in cell based assays. The crystal structure of CDK2 in complex with one of these analogues was determined and gives insight to their increased potency. The biochemical evaluation of a representative derivative is also described.
- Hoffmann-La Roche Inc., 340 Kingsland St., Nutley, NJ 07110-1199, USA.
Organizational Affiliation: