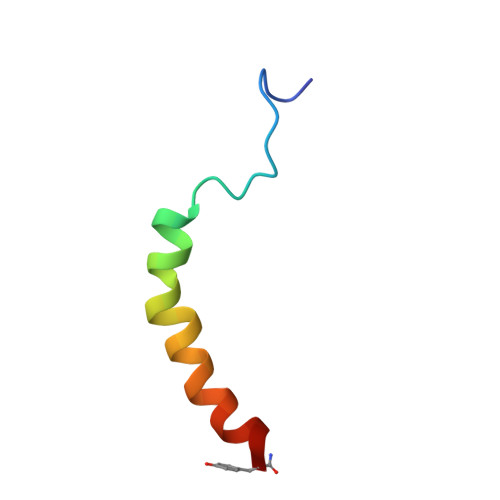
Solution structure of human neuropeptide Y.
Monks, S.A., Karagianis, G., Howlett, G.J., Norton, R.S.(1996) J Biomol NMR 8: 379-390
- PubMed: 9008359 Search on PubMed
- DOI: https://doi.org/10.1007/BF00228141
- Primary Citation Related Structures:
1RON - PubMed Abstract:
The three-dimensional structure of synthetic human neuropeptide Y in aqueous solution at pH 3.2 and 37 degrees C was determined from two-dimensional 1H NMR data recorded at 600 MHz. A restraint set consisting of 440 interproton distance restraints inferred from NOEs and 11 backbone and 4 side-chain dihedral angle restraints derived from spin-spin coupling constants was used as input for distance geometry calculations on DIANA and simulated annealing and restrained energy minimization in X-PLOR. The final set of 26 structures is well defined in the region of residues 11-36, with a mean pairwise rmsd of 0.51 A for the backbone heavy atoms (N, C alpha and C) and 1.34 A for all heavy atoms. Residues 13-36 form an amphipathic alpha-helix. The N-terminal 10 residues are poorly defined relative to the helical region, although some elements of local structure are apparent. At least one of the three prolines in the N-terminal region co-exists in both cis and trans conformations. An additional set of 24 distances was interpreted as intermolecular distances within a dimer. A combination of distance geometry and restrained simulated annealing yielded a model of the dimer having antiparallel packing of two helical units, whose hydrophobic faces form a well-defined core. Sedimentation equilibrium experiments confirm the observation that neuropeptide Y associates to form dimers and higher aggregates under the conditions of the NMR experiments. Our results therefore support the structural features reported for porcine neuropeptide Y [Cowley, D.J. et al. (1992) Eur. J. Biochem., 205, 1099-1106] rather than the 'aPP' fold described previously for human neuropeptide Y [Darbon, H. et al. (1992) Eur. J. Biochem., 209, 765-771].
- Biomolecular Research Institute, Parkville, VIC, Australia.
Organizational Affiliation: