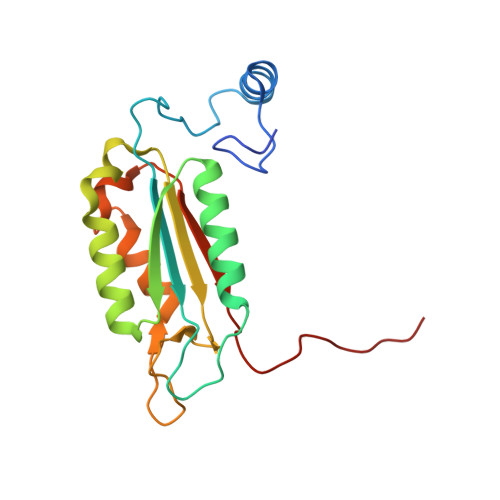
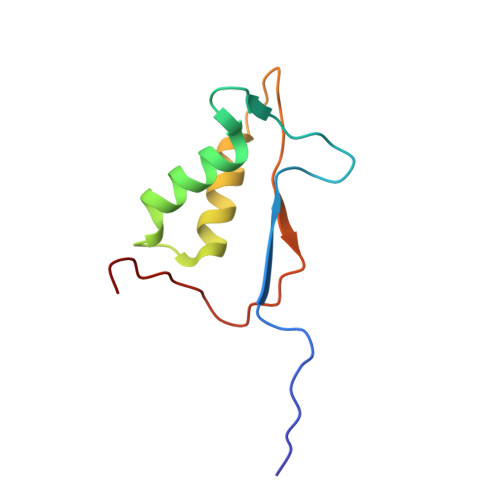
Structural analysis of caspase-1 inhibitors derived from Tethering.
O'Brien, T., Fahr, B.T., Sopko, M.M., Lam, J.W., Waal, N.D., Raimundo, B.C., Purkey, H.E., Pham, P., Romanowski, M.J.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 451-458
- PubMed: 16511067 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309105010109
- Primary Citation Related Structures:
1RWK, 1RWM, 1RWN, 1RWO, 1RWP - PubMed Abstract:
Caspase-1 is a key endopeptidase responsible for the post-translational processing of the IL-1beta and IL-18 cytokines and small-molecule inhibitors that modulate the activity of this enzyme are predicted to be important therapeutic treatments for many inflammatory diseases. A fragment-assembly approach, accompanied by structural analysis, was employed to generate caspase-1 inhibitors. With the aid of Tethering with extenders (small molecules that bind to the active-site cysteine and contain a free thiol), two novel fragments that bound to the active site and made a disulfide bond with the extender were identified by mass spectrometry. Direct linking of each fragment to the extender generated submicromolar reversible inhibitors that significantly reduced secretion of IL-1beta but not IL-6 from human peripheral blood mononuclear cells. Thus, Tethering with extenders facilitated rapid identification and synthesis of caspase-1 inhibitors with cell-based activity and subsequent structural analyses provided insights into the enzyme's ability to accommodate different inhibitor-binding modes in the active site.
- Department of Biology, Sunesis Pharmaceuticals Inc., USA.
Organizational Affiliation: