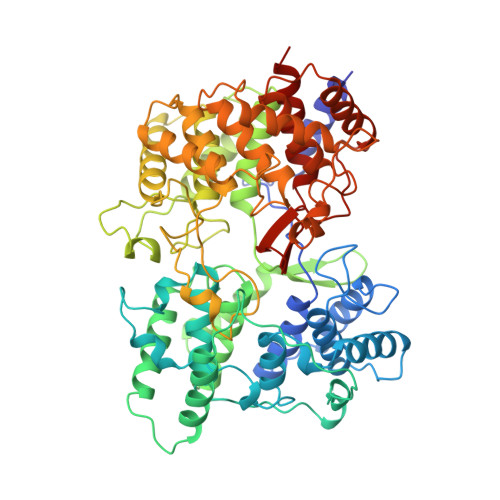
Structure and function of a squalene cyclase.
Wendt, K.U., Poralla, K., Schulz, G.E.(1997) Science 277: 1811-1815
- PubMed: 9295270
- DOI: https://doi.org/10.1126/science.277.5333.1811
- Primary Citation of Related Structures:
1SQC - PubMed Abstract:
The crystal structure of squalene-hopene cyclase from Alicyclobacillus acidocaldarius was determined at 2.9 angstrom resolution. The mechanism and sequence of this cyclase are closely related to those of 2,3-oxidosqualene cyclases that catalyze the cyclization step in cholesterol biosynthesis. The structure reveals a membrane protein with membrane-binding characteristics similar to those of prostaglandin-H2 synthase, the only other reported protein of this type. The active site of the enzyme is located in a large central cavity that is of suitable size to bind squalene in its required conformation and that is lined by aromatic residues. The structure supports a mechanism in which the acid starting the reaction by protonating a carbon-carbon double bond is an aspartate that is coupled to a histidine. Numerous surface alpha helices are connected by characteristic QW-motifs (Q is glutamine and W is tryptophan) that tighten the protein structure, possibly for absorbing the reaction energy without structural damage.
- Institut für Organische Chemie und Biochemie, Albertstrasse 21, D-79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: