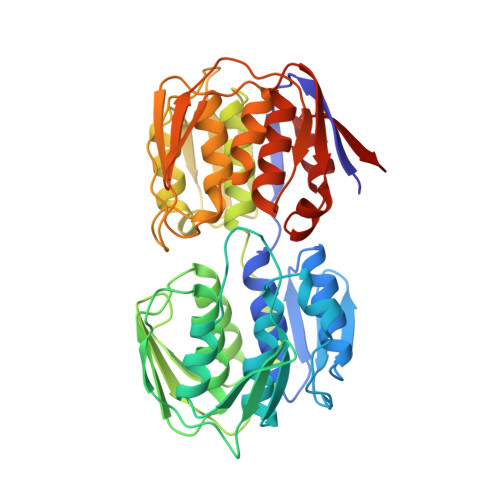
Structure of UDP-N-acetylglucosamine enolpyruvyl transferase, an enzyme essential for the synthesis of bacterial peptidoglycan, complexed with substrate UDP-N-acetylglucosamine and the drug fosfomycin.
Skarzynski, T., Mistry, A., Wonacott, A., Hutchinson, S.E., Kelly, V.A., Duncan, K.(1996) Structure 4: 1465-1474
- PubMed: 8994972 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(96)00153-0
- Primary Citation Related Structures:
1UAE - PubMed Abstract:
UDP-N-acetylglucosamine enolpyruvyl transferase (MurA), catalyses the first committed step of bacterial cell wall biosynthesis and is a target for the antibiotic fosfomycin. The only other known enolpyruvyl transferase is 5-enolpyruvylshikimate-3-phosphate (EPSP) synthase, an enzyme involved in the shikimic acid pathway and the target for the herbicide glyphosate. Inhibitors of enolpyruvyl transferases are of biotechnological interest as MurA and EPSP synthase are found exclusively in plants and microbes. The crystal structure of Escherichia coli MurA complexed with UDP-N-acetylglucosamine (UDP-GlcNAc) and fosfomycin has been determined at 1.8 A resolution. The structure consists of two domains with the active site located between them. The domains have a very similar secondary structure, and the overall protein architecture is similar to that of EPSP synthase. The fosfomycin molecule is covalently bound to the cysteine residue Cys115, whereas UDP-GlcNAc makes several hydrogen-bonding interactions with residues from both domains. The present structure reveals the mode of binding of the natural substrate UDP-GlcNAc and of the drug fosfomycin, and provides information on the residues involved in catalysis. These results should aid the design of inhibitors which would interfere with enzyme-catalyzed reactions in the early stage of the bacterial cell wall biosynthesis. Furthermore, the crystal structure of MurA provides a model for predicting active-site residues in EPSP synthase that may be involved in catalysis and substrate binding.
- Glaxo Wellcome Research and Development, Medicines Research Centre, Stevenage, UK. ts 14913@ggr.co.uk
Organizational Affiliation: