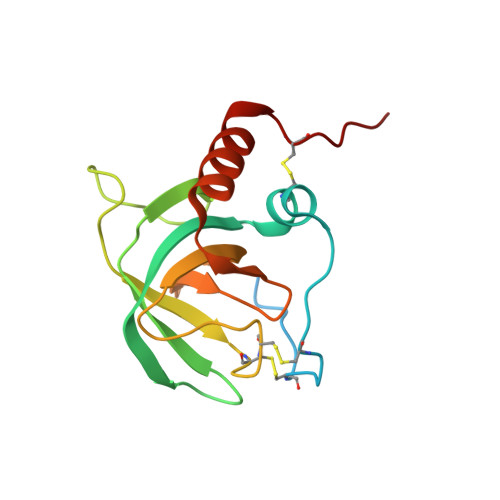
NMR Structure of the Netrin-like Domain (NTR) of Human Type I Procollagen C-Proteinase Enhancer Defines Structural Consensus of NTR Domains and Assesses Potential Proteinase Inhibitory Activity and Ligand Binding.
Liepinsh, E., Banyai, L., Pintacuda, G., Trexler, M., Patthy, L., Otting, G.(2003) J Biological Chem 278: 25982-25989
- PubMed: 12670942 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M302734200
- Primary Citation Related Structures:
1UAP - PubMed Abstract:
Procollagen C-proteinase enhancer (PCOLCE) proteins are extracellular matrix proteins that enhance the activities of procollagen C-proteinases by binding to the C-propeptide of procollagen I. PCOLCE proteins are built of three structural modules, consisting of two CUB domains followed by a C-terminal netrin-like (NTR) domain. While the enhancement of proteinase activity can be ascribed solely to the CUB domains, sequence homology of the NTR domain with tissue inhibitors of metalloproteinases suggest proteinase inhibitory activity for the NTR domain. Here we present the three-dimensional structure of the NTR domain of human PCOLCE1 as the first example of a structural domain with the canonical features of an NTR module. The structure rules out a binding mode to metalloproteinases similar to that of tissue inhibitors of metalloproteinases but suggests possible inhibitory function toward specific serine proteinases. Sequence conservation between 13 PCOLCE proteins from different organisms suggests a conserved binding surface for other protein partners.
- Department of Medical Biochemistry and Biophysics, Karolinska Institute, S-17177 Stockholm, Sweden.
Organizational Affiliation: