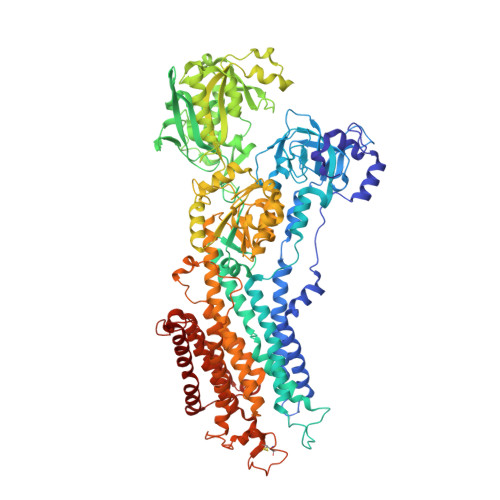
Lumenal gating mechanism revealed in calcium pump crystal structures with phosphate analogues
Toyoshima, C., Nomura, H., Tsuda, T.(2004) Nature 432: 361-368
- PubMed: 15448704 Search on PubMed
- DOI: https://doi.org/10.1038/nature02981
- Primary Citation Related Structures:
1WPG, 2ZBD - PubMed Abstract:
P-type ion transporting ATPases are ATP-powered ion pumps that establish ion concentration gradients across biological membranes. Transfer of bound cations to the lumenal or extracellular side occurs while the ATPase is phosphorylated. Here we report at 2.3 A resolution the structure of the calcium-ATPase of skeletal muscle sarcoplasmic reticulum, a representative P-type ATPase that is crystallized in the absence of Ca2+ but in the presence of magnesium fluoride, a stable phosphate analogue. This and other crystal structures determined previously provide atomic models for all four principal states in the reaction cycle. These structures show that the three cytoplasmic domains rearrange to move six out of ten transmembrane helices, thereby changing the affinity of the Ca2+-binding sites and the gating of the ion pathway. Release of ADP triggers the opening of the lumenal gate and release of phosphate its closure, effected mainly through movement of the A-domain, the actuator of transmembrane gates.
- Institute of Molecular and Cellular Biosciences, The University of Tokyo, Bunkyo-ku, Tokyo 113-0032, Japan. ct@iam.u-tokyo.ac.jp
Organizational Affiliation: