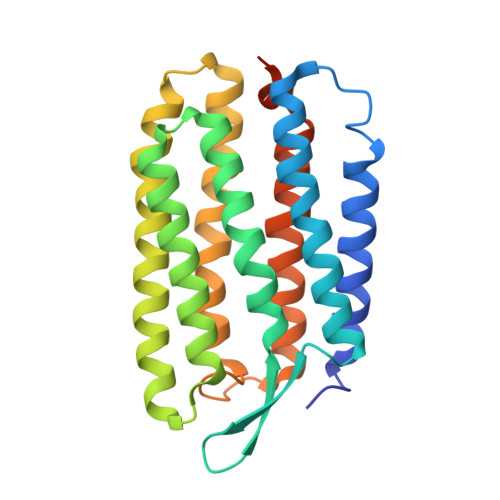
Crystal structure of the 13-cis isomer of bacteriorhodopsin in the dark-adapted state.
Nishikawa, T., Murakami, M., Kouyama, T.(2005) J Mol Biology 352: 319-328
- PubMed: 16084526 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.07.021
- Primary Citation Related Structures:
1X0S - PubMed Abstract:
The atomic structure of the trans isomer of bacteriorhodopsin was determined previously by using a 3D crystal belonging to the space group P622. Here, a structure is reported for another isomer with the 13-cis, 15-syn retinal in a dark-adapted crystal. Structural comparison of the two isomers indicates that retinal isomerization around the C13[double bond]C14 and the C15[double bond]N bonds is accompanied by noticeable displacements of a few residues in the vicinity of the retinal Schiff base and small re-arrangement of the hydrogen-bonding network in the proton release channel. On the other hand, aromatic residues surrounding the retinal polyene chain were found to scarcely move during the dark/light adaptation. This result suggests that variation in the structural rigidity within the retinal-binding pocket is one of the important factors ensuring the stereospecific isomerization of retinal.
- Department of Physics, Graduate School of Science, Nagoya University, Nagoya 464-8602, Japan.
Organizational Affiliation: