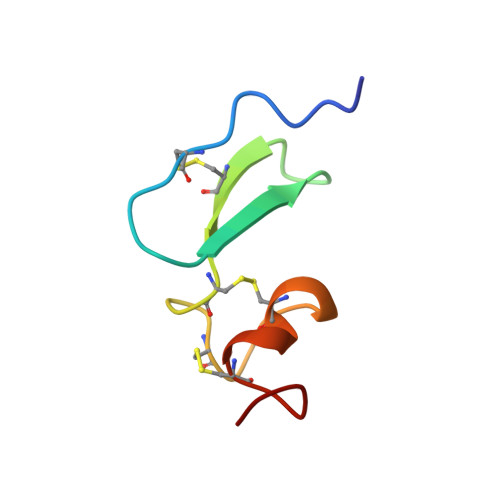
Structures of APRIL-receptor complexes: like BCMA, TACI employs only a single cysteine-rich domain for high affinity ligand binding.
Hymowitz, S.G., Patel, D.R., Wallweber, H.J., Runyon, S., Yan, M., Yin, J., Shriver, S.K., Gordon, N.C., Pan, B., Skelton, N.J., Kelley, R.F., Starovasnik, M.A.(2005) J Biological Chem 280: 7218-7227
- PubMed: 15542592 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411714200
- Primary Citation Related Structures:
1XU1, 1XU2, 1XUT - PubMed Abstract:
TACI is a member of the tumor necrosis factor receptor superfamily and serves as a key regulator of B cell function. TACI binds two ligands, APRIL and BAFF, with high affinity and contains two cysteine-rich domains (CRDs) in its extracellular region; in contrast, BCMA and BR3, the other known high affinity receptors for APRIL and BAFF, respectively, contain only a single or partial CRD. However, another form of TACI exists wherein the N-terminal CRD is removed by alternative splicing. We find that this shorter form is capable of ligand-induced cell signaling and that the second CRD alone (TACI_d2) contains full affinity for both ligands. Furthermore, we report the solution structure and alanine-scanning mutagenesis of TACI_d2 along with co-crystal structures of APRIL.TACI_d2 and APRIL.BCMA complexes that together reveal the mechanism by which TACI engages high affinity ligand binding through a single CRD, and we highlight sources of ligand-receptor specificity within the APRIL/BAFF system.
- Department of Protein Engineering, Molecular Oncology, Medicinal Chemistry, and Immunology, Genentech, Inc., South San Francisco, California 94080, USA.
Organizational Affiliation: