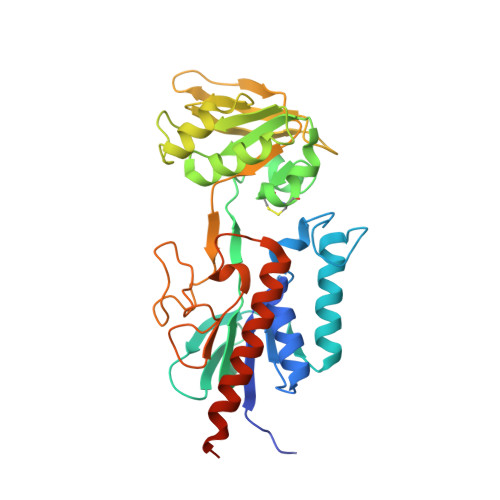
Conformational flexibility of Mycobacterium tuberculosis thioredoxin reductase: crystal structure and normal-mode analysis.
Akif, M., Suhre, K., Verma, C., Mande, S.C.(2005) Acta Crystallogr D Biol Crystallogr 61: 1603-1611
- PubMed: 16301794 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905030519
- Primary Citation Related Structures:
2A87 - PubMed Abstract:
The thioredoxin system exists ubiquitously and participates in essential antioxidant and redox-regulation processes via a pair of conserved cysteine residues. In Mycobacterium tuberculosis, which lacks a genuine glutathione system, the thioredoxin system provides reducing equivalents inside the cell. The three-dimensional structure of thioredoxin reductase from M. tuberculosis has been determined at 3 A resolution. TLS refinement reveals a large libration axis, showing that NADPH-binding domain has large anisotropic disorder. The relative rotation of the NADPH domain with respect to the FAD domain is necessary for the thioredoxin reduction cycle, as it brings the spatially distant reacting sites close together. Normal-mode analysis carried out based on the elastic network model shows that the motion required to bring about the functional conformational change can be accounted for by motion along one single mode. TLS refinement and normal-mode analysis thus enhance our understanding of the associated conformational changes.
- Centre for DNA Fingerprinting and Diagnostics, Nacharam, Hyderabad, India.
Organizational Affiliation: