Distinct Orientation of the Alloreactive Monoclonal Cd8 T Cell Activation Program by Three Different Peptide/Mhc Complexes.
Auphan-Anezin, N., Mazza, C., Guimezanes, A., Barrett-Wilt, G.A., Montero-Julian, F., Roussel, A., Hunt, D.F., Malissen, B., Schmitt-Verhulst, A.M.(2006) Eur J Immunol 36: 1856
- PubMed: 16761314 Search on PubMed
- DOI: https://doi.org/10.1002/eji.200635895
- Primary Citation Related Structures:
2CLV, 2CLZ - PubMed Abstract:
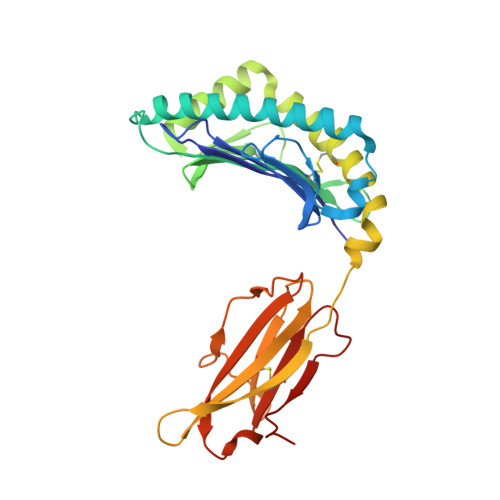
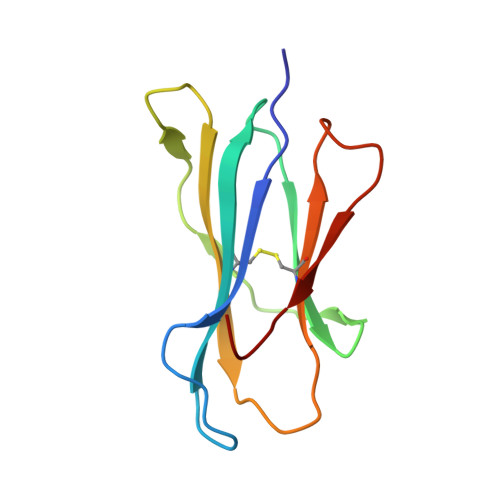
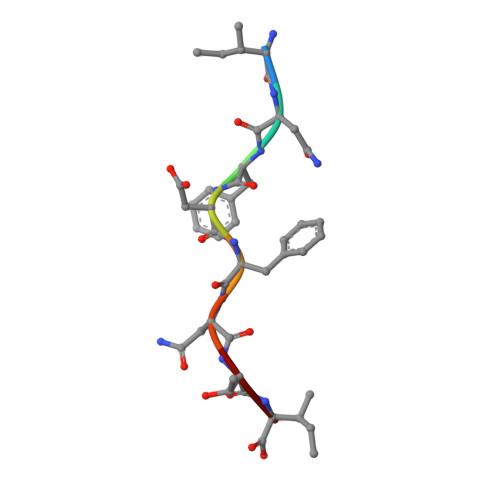
We have characterized three different programs of activation for alloreactive CD8 T cells expressing the BM3.3 TCR, their elicitation depending on the characteristics of the stimulating peptide/MHC complex. The high-affinity interaction between the TCR and the K(b)-associated endogenous peptide pBM1 (INFDFNTI) induced a complete differentiation program into effector cells correlated with sustained ERK activation. The K(bm8) variant elicited a partial activation program with delayed T cell proliferation, poor CTL activity and undetectable ERK phosphorylation; this resulted from a low-avidity interaction of TCR BM3.3 with a newly identified endogenous peptide, pBM8 (SQYYYNSL). Interestingly, mismatched pBM1/K(bm8) complexes induced a split response in BM3.3 T cells, with total reconstitution of T cell proliferation but defective generation of CTL activity that was correlated with strong but shortened ERK phosphorylation. Crystal structures highlight the molecular basis for the higher stability of pBM8/K(bm8) compared to pBM1/K(bm8) complexes that exist in two conformers. This study illustrates the importance of the stability of both peptide/MHC and peptide/MHC-TCR interactions for induction of sustained signaling required to induce optimal CTL effector functions. Subtle allelic structural variations, amplified by peptide selection, may thus orient distinct outcomes of alloreactive TCR-based therapies.
- Centre d'Immunologie de Marseille-Luminy, CNRS-INSERM-Universite de la Méditerranée, Campus de Luminy, Marseille, France. auphan@ciml.univ-mrs.fr
Organizational Affiliation: