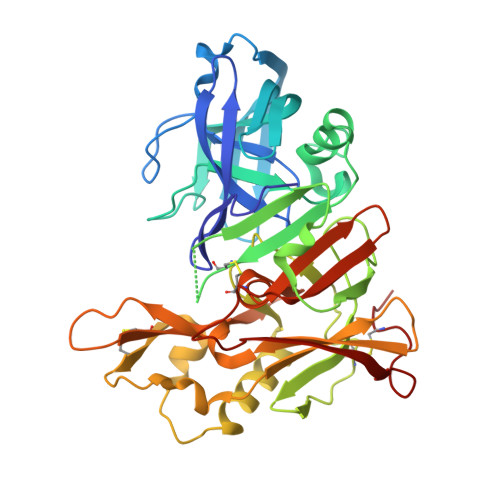
Crystal structure of human BACE2 in complex with a hydroxyethylamine transition state inhibitor
Ostermann, N., Eder, J., Eidhoff, U., Zink, F., Hassiepen, U., Worpenberg, S., Maibaum, J., Simic, O., Hommel, U., Gerhartz, B.(2006) J Mol Biology 355: 249-261
- PubMed: 16305800 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.10.027
- Primary Citation Related Structures:
2EWY - PubMed Abstract:
BACE2 is a membrane-bound aspartic protease of the A1 family with a high level of sequence homology to BACE1. While BACE1 is involved in the generation of amyloid plaques in Alzheimer's disease by cleaving Abeta-peptides from the amyloid precursor protein, the physiological function of BACE2 is not well understood. BACE2 appears to be associated with the early onset of dementia in patients with Down's syndrome, and it has been shown to be highly expressed in breast cancers. Further, it may participate in the function of normal and abnormal processes of human muscle biology. Similar to other aspartic proteases, BACE2 is expressed as an inactive zymogen requiring the cleavage of its pro-sequence during the maturation process. We have produced mature BACE2 by expression of pro-BACE2 in Escherichia coli as inclusion bodies, followed by refolding and autocatalytic activation at pH 3.4. Using a C and N-terminally truncated BACE2 variant, we were able to crystallize and determine the crystal structure of mature BACE2 in complex with a hydroxyethylamine transition-state mimetic inhibitor at 3.1 angstroms resolution. The structure of BACE2 follows the general fold of A1 aspartic proteases. However, similar to BACE1, its C-terminal domain is significantly larger than that of the other family members. Furthermore, the structure of BACE2 reveals differences in the S3, S2, S1' and S2' active site substrate pockets as compared to BACE1, and allows, therefore, for a deeper understanding of the structural features that may facilitate the design of selective BACE1 or BACE2 inhibitors.
- Protease Platform, Novartis Institutes for BioMedical Research, CH-4002 Basel, Switzerland. nils.ostermann@novartis.com
Organizational Affiliation: