STD and TRNOESY NMR studies for the epitope mapping of the phosphorylation motif of the oncogenic protein beta-catenin recognized by a selective monoclonal antibody
Megy, S., Bertho, G., Gharbi-Benarous, J., Baleux, F., Benarous, R., Girault, J.P.(2006) FEBS Lett 580: 5411-5422
- PubMed: 16996060 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2006.08.084
- Primary Citation Related Structures:
2G57 - PubMed Abstract:
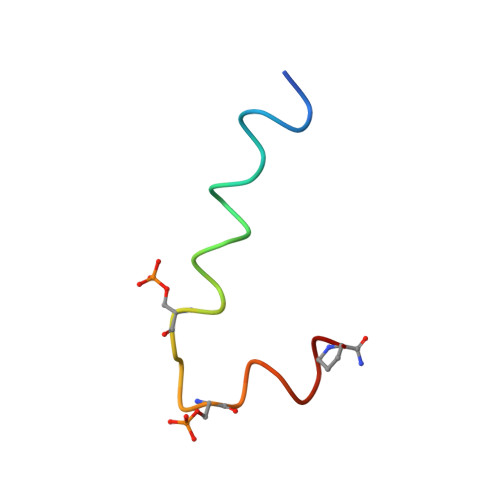
The interaction of the P-beta-Cat(19-44) peptide, a 26 amino acid peptide (K(19)AAVSHWQQQSYLDpSGIHpSGATTTAP(44)) that mimics the phosphorylated beta-Catenin antigen, has been studied with its monoclonal antibody BC-22, by transferred nuclear Overhauser effect NMR spectroscopy (TRNOESY) and saturation transfer difference NMR (STD NMR) spectroscopy. This antibody is specific to diphosphorylated beta-Catenin and does not react with the non-phosphorylated protein. Phosphorylation of beta-Catenin at sites Ser33 and Ser37 on the DSGXXS motif is required for the interaction of beta-Catenin with the ubiquitin ligase SCF(beta-TrCP). beta-TrCP is involved in the ubiquitination and proteasome targeting of the oncogenic protein beta-Catenin, the accumulation of which has been implicated in various human cancers. The three-dimensional structure of the P-beta-Cat(19-44) in the bound conformation was determined by TRNOESY NMR experiments; the peptide adopts a compact structure in the presence of mAb with formation of turns around Trp25 and Gln26, with a tight bend created by the DpS(33)GIHpS(37) motif; the peptide residues (D32-pS37) forming this bend are recognized by the antibody as demonstrated by STD NMR experiments. STD NMR studies provide evidence for the existence of a conformational epitope containing tandem repeats of phosphoserine motifs. The peptide's epitope is predominantly located in the large bend and in the N-terminal segment, implicating bidentate association. These findings are in excellent agreement with a recently published NMR structure required for the interaction of beta-Catenin with the SCF(beta-TrCP) protein.
- Université Paris V-René Descartes, Laboratoire de Chimie et Biochimie Pharmacologiques et Toxicologiques, UMR 8601 CNRS, 45 Rue des Saint-Pères, 75270 Paris Cedex 06, France.
Organizational Affiliation: