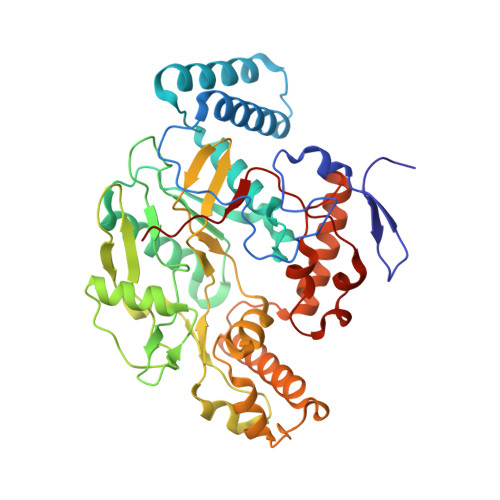
Structural studies of constitutive nitric oxide synthases with diatomic ligands bound.
Li, H., Igarashi, J., Jamal, J., Yang, W., Poulos, T.L.(2006) J Biol Inorg Chem 11: 753-768
- PubMed: 16804678 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-006-0123-8
- Primary Citation Related Structures:
2G6H, 2G6I, 2G6J, 2G6K, 2G6L, 2G6M, 2G6N, 2G6O - PubMed Abstract:
Crystal structures are reported for the endothelial nitric oxide synthase (eNOS)-arginine-CO ternary complex as well as the neuronal nitric oxide synthase (nNOS) heme domain complexed with L: -arginine and diatomic ligands, CO or NO, in the presence of the native cofactor, tetrahydrobiopterin, or its oxidized analogs, dihydrobiopterin and 4-aminobiopterin. The nature of the biopterin has no influence on the diatomic ligand binding. The binding geometries of diatomic ligands to nitric oxide synthase (NOS) follow the {MXY}(n) formalism developed from the inorganic diatomic-metal complexes. The structures reveal some subtle structural differences between eNOS and nNOS when CO is bound to the heme which correlate well with the differences in CO stretching frequencies observed by resonance Raman techniques. The detailed hydrogen-bonding geometries depicted in the active site of nNOS structures indicate that it is the ordered active-site water molecule rather than the substrate itself that would most likely serve as a direct proton donor to the diatomic ligands (CO, NO, as well as O(2)) bound to the heme. This has important implications for the oxygen activation mechanism critical to NOS catalysis.
- Department of Molecular Biology and Biochemistry, Center in Chemical and Structural Biology, University of California, Irvine, CA 92697-3900, USA.
Organizational Affiliation: