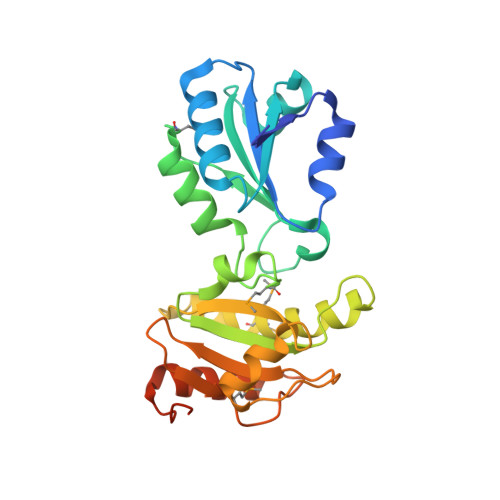
Crystal Structure of the bb' Domains of the Protein Disulfide Isomerase ERp57.
Kozlov, G., Maattanen, P., Schrag, J.D., Pollock, S., Cygler, M., Nagar, B., Thomas, D.Y., Gehring, K.(2006) Structure 14: 1331-1339
- PubMed: 16905107 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2006.06.019
- Primary Citation Related Structures:
2H8L - PubMed Abstract:
The synthesis of proteins in the endoplasmic reticulum (ER) is limited by the rate of correct disulfide bond formation. This process is carried out by protein disulfide isomerases, a family of ER proteins which includes general enzymes such as PDI that recognize unfolded proteins and others that are selective for specific proteins or classes. Using small-angle X-ray scattering and X-ray crystallography, we report the structure of a selective isomerase, ERp57, and its interactions with the lectin chaperone calnexin. Using isothermal titration calorimetry and NMR spectroscopy, we show that the b' domain of ERp57 binds calnexin with micromolar affinity through a conserved patch of basic residues. Disruption of this binding site by mutagenesis abrogates folding of RNase B in an in vitro assay. The relative positions of the ERp57 catalytic sites and calnexin binding site suggest that activation by calnexin is due to substrate recruitment rather than a direct stimulation of ERp57 oxidoreductase activity.
- Biochemistry Department, McGill University, 3655 Promenade Sir William Osler, Montréal, Québec H3G 1Y6, Canada.
Organizational Affiliation: