The solution structure of the copper(I)-mediated complex between the first soluble domain of the Menkes protein and the metallochaperone HAH1.
Bertini, I., Banci, L.C., Felli, I.C., Pavelkova, A., Rosato, A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Copper-transporting ATPase 1 | 73 | Homo sapiens | Mutation(s): 0 Gene Names: ATP7A, MC1, MNK EC: 3.6.3.4 (PDB Primary Data), 7.2.2.8 (UniProt) | 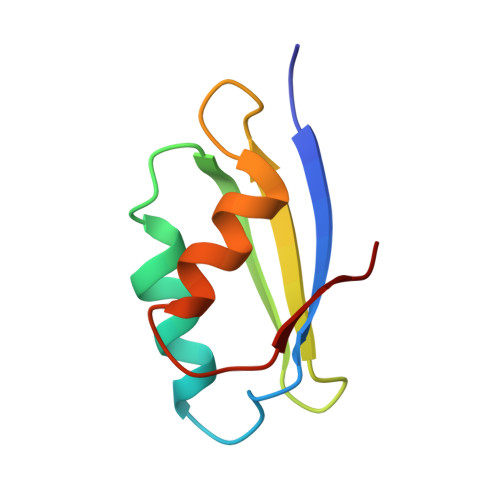 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q04656 GTEx: ENSG00000165240 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q04656 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Copper transport protein ATOX1 | 68 | Homo sapiens | Mutation(s): 0 Gene Names: ATOX1, HAH1 | 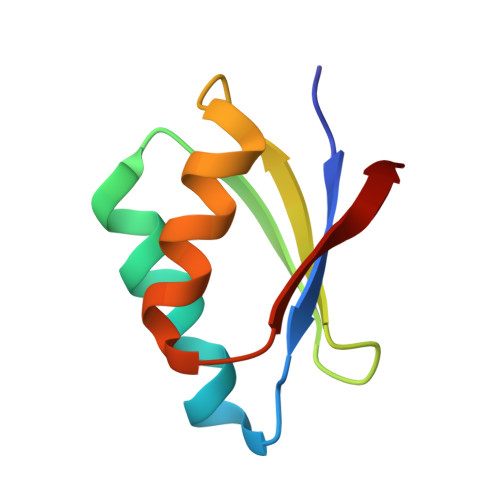 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O00244 GTEx: ENSG00000177556 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O00244 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CU Download:Ideal Coordinates CCD File | C [auth A] | COPPER (II) ION Cu JPVYNHNXODAKFH-UHFFFAOYSA-N |  | ||