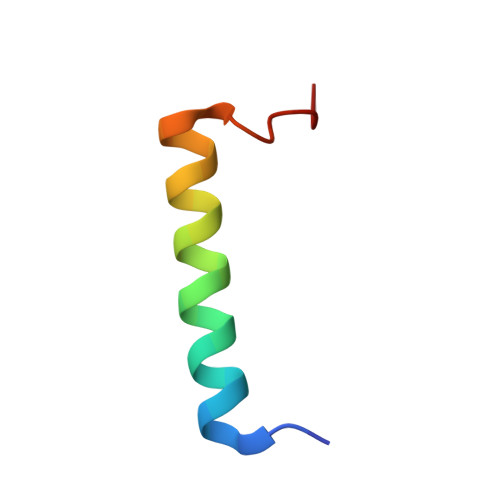
Solution NMR Structure of Apo-Calmodulin in Complex with the IQ Motif of Human Cardiac Sodium Channel NaV1.5.
Chagot, B., Chazin, W.J.(2011) J Mol Biology 406: 106-119
- PubMed: 21167176 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.11.046
- Primary Citation Related Structures:
2L53 - PubMed Abstract:
The function of the human voltage-gated sodium channel Na(V)1.5 is regulated in part by intracellular calcium signals. The ubiquitous calcium sensor protein calmodulin (CaM) is an important part of the complex calcium-sensing apparatus in Na(V)1.5. CaM interacts with an IQ (isoleucine-glutamine) motif in the large intracellular C-terminal domain of the channel. Using co-expression and co-purification, we have been able to isolate a CaM-IQ motif complex and to determine its high-resolution structure in absence of calcium using multi-dimensional solution NMR. Under these conditions, the Na(V)1.5 IQ motif interacts with the C-terminal domain (C-lobe) of CaM, with the N-terminal domain remaining free in solution. The structure reveals that the C-lobe adopts a semi-open conformation with the IQ motif bound in a narrow hydrophobic groove. Sequence similarities between voltage-gated sodium channels and voltage-gated calcium channels suggest that the structure of the CaM-Na(V)1.5 IQ motif complex can serve as a general model for the interaction between CaM and ion channel IQ motifs under low-calcium conditions. The structure also provides insight into the biochemical basis for disease-associated mutations that map to the IQ motif in Na(V)1.5.
- Department of Biochemistry, Vanderbilt University, Nashville, TN 37232, USA.
Organizational Affiliation: