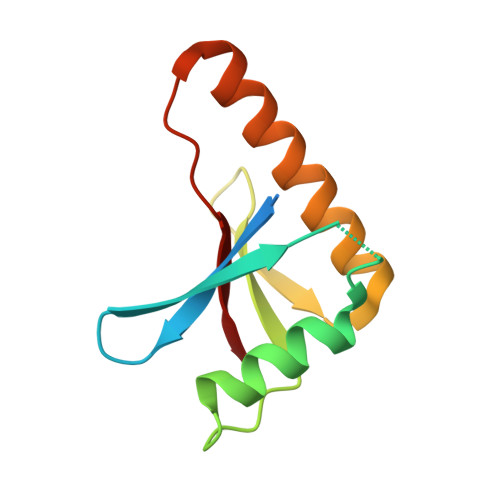
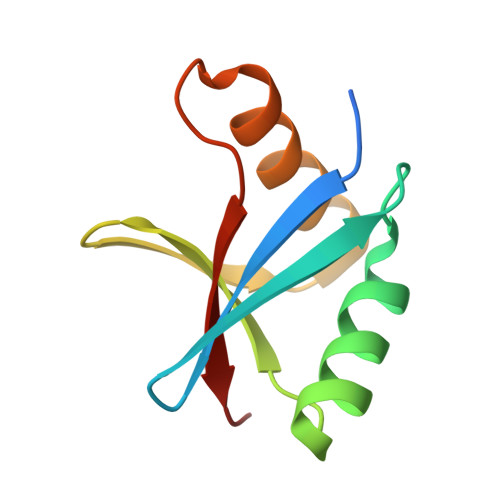
Crystal Structure of the Complex of Human Mitogen Activated Protein Kinase Kinase 5 Phox Domain (MAP2K5-phox) with Human Mitogen Activated Protein Kinase Kinase Kinase 3 (MAP3K3B-phox)
Filippakopoulos, P., Savitsky, P., Ugochukwu, E., Edwards, A., Arrowsmith, C., Sundstrom, M., von Delft, F., Knapp, S.To be published.