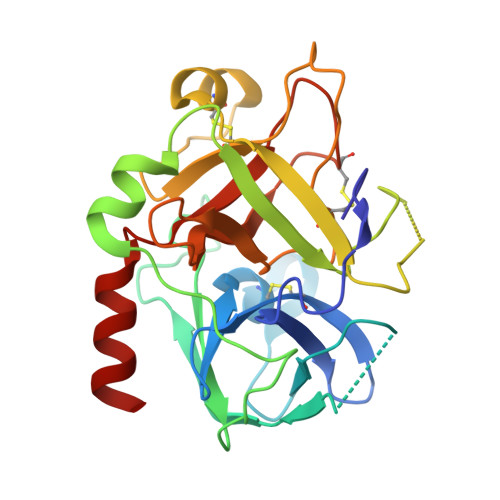
Structural basis of na+ activation mimicry in murine thrombin.
Marino, F., Chen, Z.W., Ergenekan, C.E., Bush-Pelc, L.A., Mathews, F.S., Di Cera, E.(2007) J Biological Chem 282: 16355-16361
- PubMed: 17428793 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M701323200
- Primary Citation Related Structures:
2OCV, 2OD3 - PubMed Abstract:
Unlike human thrombin, murine thrombin lacks Na+ activation due to the charge reversal substitution D222K in the Na+ binding loop. However, the enzyme is functionally stabilized in a Na+-bound form and is highly active toward physiologic substrates. The structural basis of this peculiar property is unknown. Here, we present the 2.2 A resolution x-ray crystal structure of murine thrombin in the absence of inhibitors and salts. The enzyme assumes an active conformation, with Ser-195, Glu-192, and Asp-189 oriented as in the Na+-bound fast form of human thrombin. Lys-222 completely occludes the pore of entry to the Na+ binding site and positions its side chain inside the pore, with the Nzeta atom H-bonded to the backbone oxygen atoms of Lys-185, Asp-186b, and Lys-186d. The same architecture is observed in the 1.75 A resolution structure of a thrombin chimera in which the human enzyme carries all residues defining the Na+ pore in the murine enzyme. These findings demonstrate that Na+ activation in thrombin is linked to the architecture of the Na+ pore. The molecular strategy of Na+ activation mimicry unraveled for murine thrombin is relevant to serine proteases and enzymes activated by monovalent cations in general.
- Department of Biochemistry and Molecular Biophysics, Washington University Medical School, St. Louis, Missouri 63110, USA.
Organizational Affiliation: