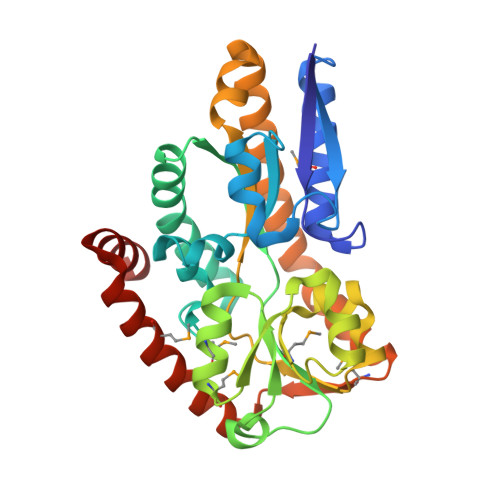
Crystal Structures of two Bordetella pertussis Periplasmic Receptors Contribute to Defining a Novel Pyroglutamic Acid Binding DctP Subfamily.
Rucktooa, P., Antoine, R., Herrou, J., Huvent, I., Locht, C., Jacob-Dubuisson, F., Villeret, V., Bompard, C.(2007) J Mol Biology 370: 93-106
- PubMed: 17499270 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.04.047
- Primary Citation Related Structures:
2PFY, 2PFZ - PubMed Abstract:
Gram-negative bacteria have developed several different transport systems for solute uptake. One of these, the tripartite ATP independent periplasmic transport system (TRAP-T), makes use of an extracytoplasmic solute receptor (ESR) which captures specific solutes with high affinity and transfers them to their partner permease complex located in the bacterial inner membrane. We hereby report the structures of DctP6 and DctP7, two such ESRs from Bordetella pertussis. These two proteins display a high degree of sequence and structural similarity and possess the "Venus flytrap" fold characteristic of ESRs, comprising two globular alpha/beta domains hinged together to form a ligand binding cleft. DctP6 and DctP7 both show a closed conformation due to the presence of one pyroglutamic acid molecule bound by highly conserved residues in their respective ligand binding sites. BLAST analyses have revealed that the DctP6 and DctP7 residues involved in ligand binding are strictly present in a number of predicted TRAP-T ESRs from other bacteria. In most cases, the genes encoding these TRAP-T systems are located in the vicinity of a gene coding for a pyroglutamic acid metabolising enzyme. Both the high degree of conservation of these ligand binding residues and the genomic context of these TRAP-T-coding operons in a number of bacterial species, suggest that DctP6 and DctP7 constitute the prototypes of a novel TRAP-T DctP subfamily involved in pyroglutamic acid transport.
- UMR 8161 CNRS Institut de Biologie de Lille, Laboratoire de Cristallographie Macromoléculaire, Université des Sciences et Technologies de Lille, Université de Lille 2, Institut Pasteur de Lille IFR142, Lille cedex, France.
Organizational Affiliation: