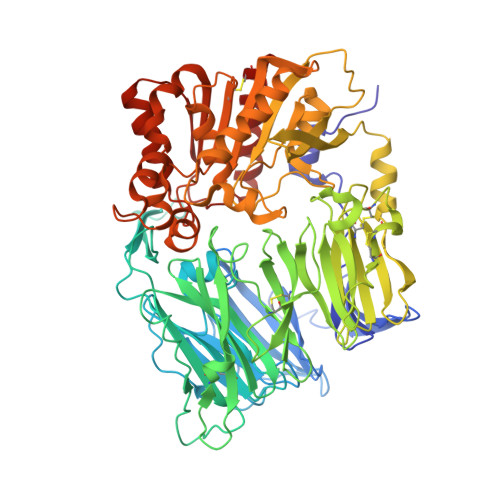
Synthesis, SAR, and X-ray structure of novel potent DPPIV inhibitors: oxadiazolyl ketones.
Koo, K.D., Kim, M.J., Kim, S., Kim, K.H., Hong, S.Y., Hur, G.C., Yim, H.J., Kim, G.T., Han, H.O., Kwon, O.H., Kwon, T.S., Koh, J.S., Lee, C.S.(2007) Bioorg Med Chem Lett 17: 4167-4172
- PubMed: 17544668 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.05.046
- Primary Citation Related Structures:
2QKY - PubMed Abstract:
Synthesis of a novel series of DPPIV inhibitors with 1,2,4- and 1,3,4-oxadiazolyl ketone derivatives and its structure-activity relationships are discussed. Compound 18h showed good inhibitory activity against DPPIV and favorable pharmacokinetic properties. In vivo pharmacodynamic efficacy and co-crystal structure of compound 18h with DPPIV is also described.
- LG Life Sciences, Ltd/R&D Park, 104-1 Munji-dong, Yuseong-gu, Daejeon 305-380, Republic of Korea.
Organizational Affiliation: