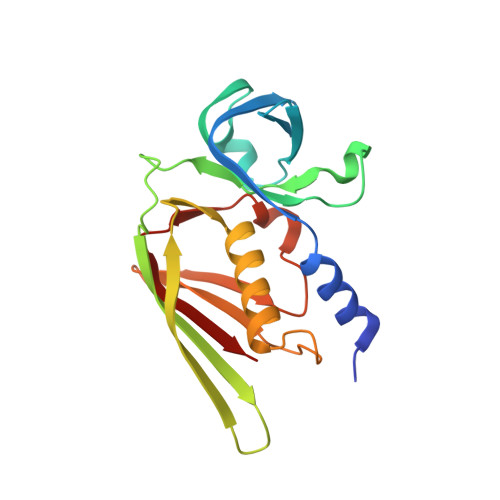
The crystal structure of staphylococcal superantigen-like protein 11 in complex with sialyl Lewis X reveals the mechanism for cell binding and immune inhibition
Chung, M.C., Wines, B.D., Baker, H., Langley, R.J., Baker, E.N., Fraser, J.D.(2007) Mol Microbiol 66: 1342-1355
- PubMed: 18045383 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2007.05989.x
- Primary Citation Related Structures:
2RDG, 2RDH - PubMed Abstract:
Staphylococcus aureus is a major pathogen that produces a family of 14 staphylococcal superantigen-like (SSL) proteins, which are structurally similar to superantigens but do not stimulate T cells. SSL11 is one member of the family that is found in all staphylococcal strains. Recombinant SSL11 bound to granulocytes and monocytes through a sialic acid-dependent mechanism and was rapidly internalized. SSL11 also bound to sialic acid-containing glycoproteins, such as the Fc receptor for IgA (FcalphaRI) and P-selectin glycoprotein ligand-1 (PSGL-1), and inhibited neutrophil attachment to a P-selectin-coated surface. Biosensor analysis of two SSL11 alleles binding to sialyl Lewis X [sLe(x)- Neu5Acalpha2-3Galbeta1-4(Fuc1-3)GlcNAc] coupled to bovine serum albumin gave dissociation constants of 0.7 and 7 mum respectively. Binding of SSL11 to a glycan array revealed specificity for glycans containing the trisaccharide sialyllactosamine (sLacNac - Neu5Acalpha2-3Galbeta1-4GlcNAc). A 1.6 A resolution crystal structure of SSL11 complexed with sLe(x) revealed a discrete binding site in the C-terminal beta-grasp domain, with predominant interactions with the sialic acid and galactose residues. A single amino acid mutation in the carbohydrate binding site abolished all SSL11 binding. Thus, SSL11 is a staphylococcal protein that targets myeloid cells by binding sialyllactosamine-containing glycoproteins.
- Maurice Wilkins Centre, University of Auckland, Auckland, New Zealand.
Organizational Affiliation: