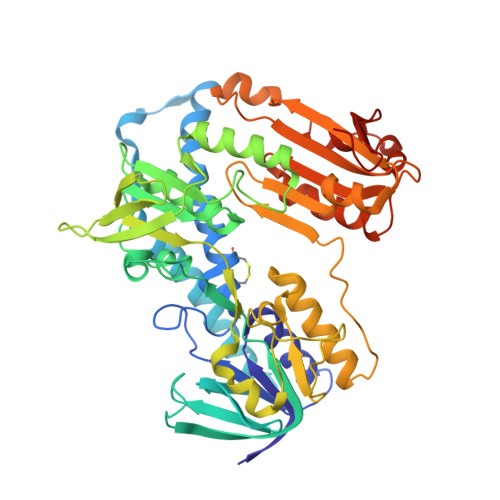
X-ray structure of trypanothione reductase from Crithidia fasciculata at 2.4-A resolution.
Kuriyan, J., Kong, X.P., Krishna, T.S., Sweet, R.M., Murgolo, N.J., Field, H., Cerami, A., Henderson, G.B.(1991) Proc Natl Acad Sci U S A 88: 8764-8768
- PubMed: 1924336
- DOI: https://doi.org/10.1073/pnas.88.19.8764
- Primary Citation of Related Structures:
2TPR - PubMed Abstract:
Trypanosomes and related protozoan parasites lack glutathione reductase and possess instead a closely related enzyme that serves as the reductant of a bis(glutathione)-spermidine conjugate, trypanothione. The human and parasite enzymes have mutually exclusive substrate specificities, providing a route for the design of therapeutic agents by specific inhibition of the parasite enzyme. We report here the three-dimensional structure of trypanothione reductase from Crithidia fasciculata and show that it closely resembles the structure of human glutathione reductase. In particular, the core structure surrounding the catalytic machinery is almost identical in the two enzymes. However, significant differences are found at the substrate binding sites. A cluster of basic residues in glutathione reductase is replaced by neutral, hydrophobic, or acidic residues in trypanothione reductase, consistent with the nature of the spermidine linkage and the change in overall charge of the substrate from -2 to +1, respectively. The binding site is more open in trypanothione reductase due to rotations of about 4 degrees in the domains that form the site, with relative shifts of as much as 2-3 A in residue positions. These results provide a detailed view of the residues that can interact with potential inhibitors and complement previous modeling and mutagenesis studies on the two enzymes.
- Howard Hughes Medical Institute, Rockefeller University, New York, NY 10021.
Organizational Affiliation: