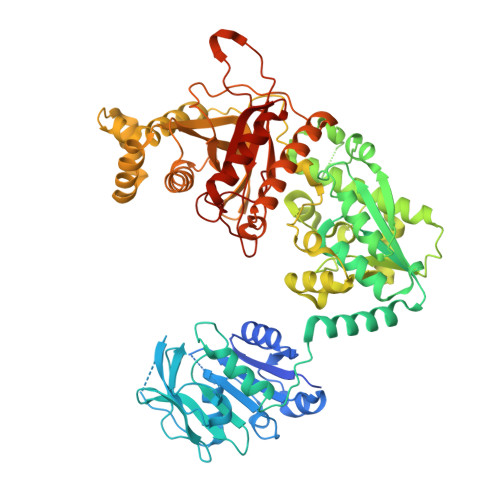
Substrate Specificity and Oligomerization of Human Gmp Synthetase
Welin, M., Lehtio, L., Johansson, A., Flodin, S., Nyman, T., Tresaugues, L., Hammarstrom, M., Graslund, S., Nordlund, P.(2013) J Mol Biology 425: 4323
- PubMed: 23816837
- DOI: https://doi.org/10.1016/j.jmb.2013.06.032
- Primary Citation of Related Structures:
2VXO - PubMed Abstract:
Guanine monophosphate (GMP) synthetase is a bifunctional two-domain enzyme. The N-terminal glutaminase domain generates ammonia from glutamine and the C-terminal synthetase domain aminates xanthine monophosphate (XMP) to form GMP. Mammalian GMP synthetases (GMPSs) contain a 130-residue-long insert in the synthetase domain in comparison to bacterial proteins. We report here the structure of a eukaryotic GMPS. Substrate XMP was bound in the crystal structure of the human GMPS enzyme. XMP is bound to the synthetase domain and covered by a LID motif. The enzyme forms a dimer in the crystal structure with subunit orientations entirely different from the bacterial counterparts. The inserted sub-domain is shown to be involved in substrate binding and dimerization. Furthermore, the structural basis for XMP recognition is revealed as well as a potential allosteric site. Enzymes in the nucleotide metabolism typically display an increased activity in proliferating cells due to the increased need for nucleotides. Many drugs used as immunosuppressants and for treatment of cancer and viral diseases are indeed nucleobase- and nucleoside-based compounds, which are acting on or are activated by enzymes in this pathway. The information obtained from the crystal structure of human GMPS might therefore aid in understanding interactions of nucleoside-based drugs with GMPS and in structure-based design of GMPS-specific inhibitors.
- Structural Genomics Consortium, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-17177 Stockholm, Sweden.
Organizational Affiliation: