Design, Synthesis and Evaluation of Indolinones as Inhibitors of the Transforming Growth Factor Beta Receptor I (Tgfbri)
Roth, G.J., Heckel, A., Brandl, T., Grauert, M., Hoerer, S., Kley, J.T., Schnapp, G., Baum, P., Mennerich, D., Schnapp, A., Park, J.E.(2010) J Med Chem 53: 7287
- PubMed: 20919678 Search on PubMed
- DOI: https://doi.org/10.1021/jm100812a
- Primary Citation Related Structures:
2X7O - PubMed Abstract:
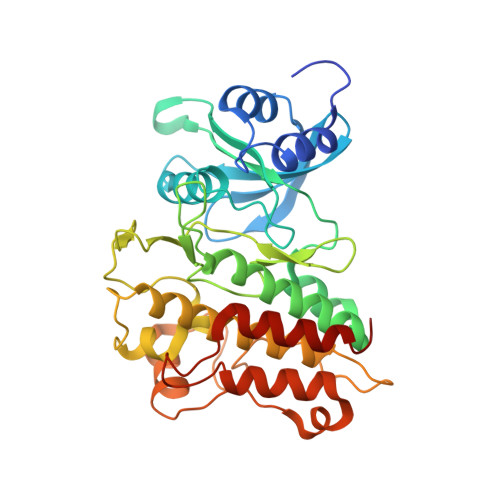
Inhibition of transforming growth factor β (TGFβ) type I receptor (Alk5) offers a novel approach for the treatment of fibrotic diseases and cancer. Indolinones substituted in position 6 were identified as a new chemotype inhibiting TGFβRI concomitant with a low cross-reactivity among the human kinome. A subset of compounds showed additional inhibition of platelet-derived growth factor receptor alpha (PDGFRα), contributing to an interesting pharmacological profile. In contrast, p38 kinase, which is often inhibited by TGFβRI inhibitors, was not targeted by derivatives based on the indolinone chemotype. Guided by an X-ray structure of lead compound 5 (BIBF0775) soaked into the kinase domain of TGFβRI, optimization furnished potent and selective inhibitors of TGFβRI. Potent inhibition translated well into good inhibition of TGFβRI-mediated phosphorylation of Smad2/3, demonstrating efficacy in a cellular setting. Optimized compounds were extensively profiled on a 232-kinase panel and showed low cross-reactivities within the human kinome.
- Biberach, Department of Medicinal Chemistry, Boehringer Ingelheim Pharma GmbH & Co KG, Birkendorfer Strasse 65,D-88397 Biberach, Germany. gerald.roth@boehringer-ingelheim.com
Organizational Affiliation: