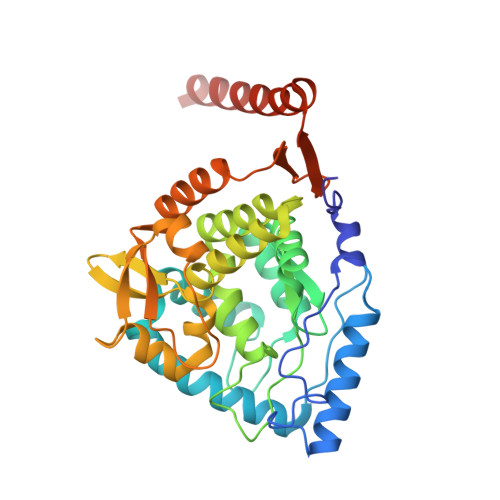
Crystal Structure of Human Tyrosine Hydroxylase Catalytic Domain
Muniz, J.R.C., Cooper, C.D.O., Yue, W.W., Krysztofinska, E., von Delft, F., Knapp, S., Gileadi, O., Arrowsmith, C.H., Edwards, A.M., Weigelt, J., Bountra, C., Kavanagh, K.L., Oppermann, U.To be published.